Reviving Lost Potential: Strategies to Recapitulate Bioactivity in Isolated Natural Compounds
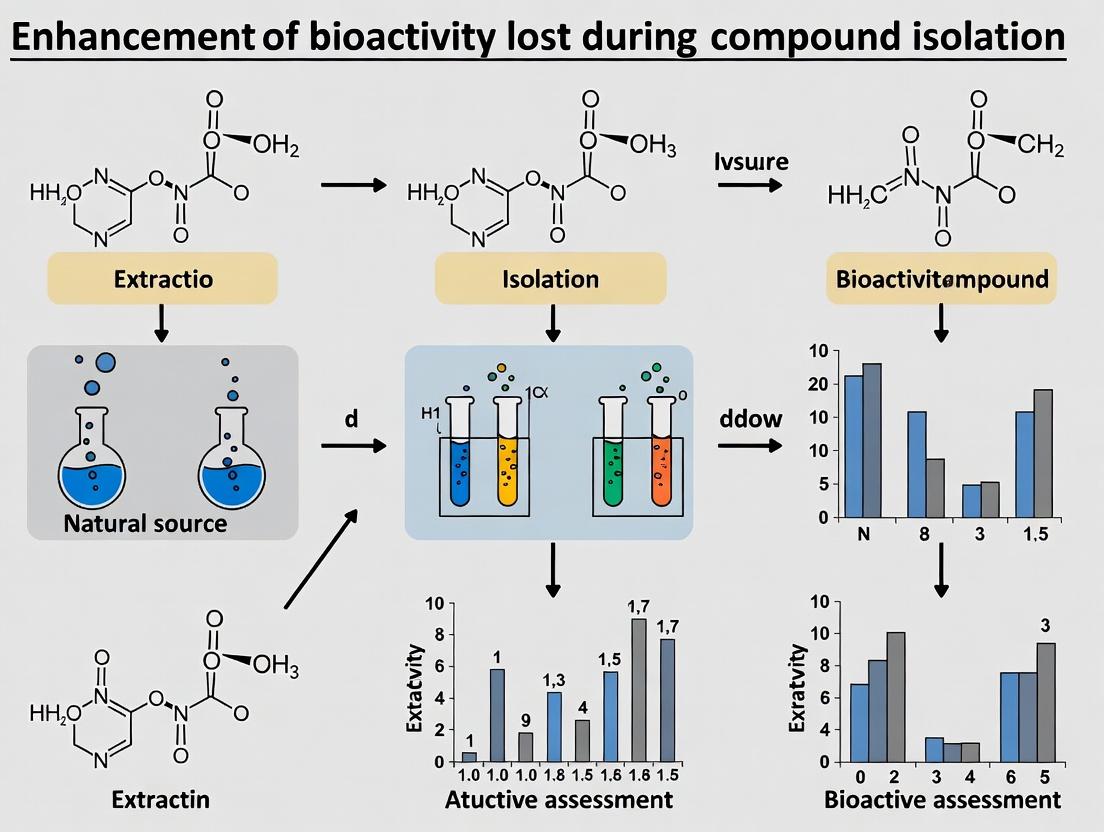
This article addresses the critical challenge of bioactivity loss observed during the isolation of compounds from natural sources, biological extracts, or complex mixtures.
Reviving Lost Potential: Strategies to Recapitulate Bioactivity in Isolated Natural Compounds
Abstract
This article addresses the critical challenge of bioactivity loss observed during the isolation of compounds from natural sources, biological extracts, or complex mixtures. Targeted at researchers and drug development professionals, it explores the underlying causes—from the disruption of synergistic networks to altered physicochemical properties—and presents a comprehensive methodological toolkit to counteract this phenomenon. We detail advanced techniques for bioactivity-guided fractionation, in situ reconstitution, and the use of adjuvant systems. The piece further provides troubleshooting frameworks for optimizing isolation protocols and discusses robust validation strategies, including phenotypic and target-based assays, to confirm the successful recapitulation of therapeutic effects. This guide synthesizes current research to empower scientists in maximizing the translational potential of bioactive discoveries.
The Bioactivity Conundrum: Why Isolated Compounds Lose Their Therapeutic Punch
Welcome to the Bioactivity Troubleshooting & Support Center. This resource is designed to help researchers diagnose and address the critical challenge of bioactivity loss during compound isolation—a major bottleneck in natural product drug discovery and lead development.
FAQs & Troubleshooting Guides
Q1: My crude extract shows potent inhibitory activity in a cell-based assay, but the purified compound is inactive. What are the most common causes?
- A: This is the core phenomenon. Primary causes include:
- Synergistic Effects Lost: Bioactivity in the crude extract results from multiple compounds acting together. Isolation removes essential co-factors.
- Compound Instability: The purified molecule degrades under isolation conditions (pH, light, temperature) or during storage.
- Altered Solubility/Bioavailability: The purified compound may not solubilize effectively in the assay buffer, unlike in the crude mixture where other components act as natural solubilizers.
- Incorrect Structural Identification: The isolated compound may be an artifact or an inactive analog of the true active principle.
- Assay Sensitivity: The concentration of the single compound in the assay is now below the effective threshold.
Q2: I suspect synergy is the issue. How can I experimentally test for this post-isolation?
- A: Implement a Fraction Recombination Assay.
- Protocol: 1) Fractionate your crude extract (e.g., via HPLC). 2) Test all individual fractions for bioactivity. 3) Systematically recombine inactive fractions in pairs or groups. 4) Re-test the combinations for restored activity. This pinpoints which fractions contain necessary cooperative elements.
Q3: How can I stabilize a seemingly labile purified compound during and after isolation?
- A: Stability must be proactively managed.
- Protocol for Stability Assessment: 1) Prepare aliquots of the purified compound in different conditions: various buffers (pH 4, 7, 10), under light vs. dark, at 4°C, -20°C, and -80°C. 2) Analyze aliquots by LC-MS at time zero (T0), after 24h, 7 days, and 30 days. 3) Monitor for decreases in parent compound peak area and the appearance of new degradation peaks. 4) Stabilize based on results: add antioxidants (e.g., ascorbic acid), chelating agents (e.g., EDTA), use amber vials, and establish a stable storage format (e.g., lyophilized powder under inert gas).
Q4: My isolated compound is poorly soluble in aqueous assay buffers. What can I do?
- A: Employ bio-compatible solubilization strategies.
- Protocol for Solubilization Optimization: Start with a stock solution in a minimal amount of DMSO (typically ≤0.5% final assay concentration). If precipitation occurs, consider: 1) Co-solvents: Add a low percentage of ethanol or PEG-400 to the buffer. 2) Complexing Agents: Use cyclodextrins (e.g., HP-β-CD) to form water-soluble inclusion complexes. 3) Delivery Vehicles: Use liposomes or albumin to shuttle the compound. Critical Control: Include the vehicle at the same concentration in all assay controls.
Table 1: Common Causes of Bioactivity Loss and Their Estimated Frequency
| Cause of Loss | Estimated Frequency* | Key Diagnostic Test |
|---|---|---|
| Synergy/Co-factor Loss | 40-50% | Fraction Recombination Assay |
| Compound Instability | 25-35% | LC-MS Stability Profiling |
| Solubility/Bioavailability Issues | 15-25% | Dynamic Light Scattering, Microscopy |
| Incorrect Identification | 5-10% | Re-isolation & Orthogonal NMR/HR-MS |
| Assay-Related Issues | 5-10% | Dose-Response with Crude Extract Standard |
Frequency estimates based on literature reviews of natural product isolation studies.
Table 2: Efficacy of Common Stabilization Agents
| Stabilizing Agent | Target Issue | Recommended Final Conc. | Typical Efficacy Increase* |
|---|---|---|---|
| Ascorbic Acid | Oxidation | 0.1-1.0 mM | 2-5 fold half-life |
| EDTA (Disodium) | Metal-Catalyzed Degradation | 0.1-0.5 mM | 3-8 fold half-life |
| HP-β-Cyclodextrin | Aqueous Solubility | 1-5 mM | 10-50 fold solubility ↑ |
| Lyophilization (at -80°C) | General Long-Term Storage | N/A | 12-24 month stability |
Efficacy is compound-dependent; these are generalized ranges.
Experimental Protocols
Protocol 1: Comprehensive Fraction Recombination Assay Objective: To identify synergistic interactions between fractions of a bioactive crude extract.
- Fractionation: Separate 100 mg of bioactive crude extract using reversed-phase flash chromatography (e.g., C18 column, gradient 5-100% MeCN in H₂O + 0.1% FA).
- Primary Screening: Test each fraction (evaporated, re-dissolved in DMSO/assay buffer) in the primary bioassay at a concentration equivalent to the original crude extract.
- Recombination: Pool inactive fractions in a systematic matrix (e.g., binary combinations). Evaporate and re-dissolve pools.
- Secondary Screening: Re-test all pools in the bioassay. Active pools indicate synergistic fractions.
- Deconvolution: Further fractionate active pools and repeat the process to narrow down the synergistic partner compounds.
Protocol 2: LC-MS Stability Profiling Workflow Objective: To quantify the degradation kinetics of a purified compound under various conditions.
- Sample Preparation: Dissolve purified compound to 1 mM in primary solvent (e.g., DMSO). Dilute 1:100 into the following stress buffers: PBS (pH 7.4), Acetate buffer (pH 5.0), Carbonate buffer (pH 9.0). Aliquot 100 µL into microtubes.
- Stress Conditions: Incubate aliquots at: 4°C (dark), 25°C (light), 25°C (dark), 40°C (dark).
- Time-Point Sampling: Withdraw samples at T=0, 1h, 6h, 24h, 7d. Immediately quench by placing on dry ice, then store at -80°C until analysis.
- LC-MS Analysis: Use a standardized LC-MS method. Integrate peak areas for the parent compound and any new peaks.
- Data Analysis: Plot % parent compound remaining vs. time for each condition. Calculate degradation rate constants and half-lives.
Pathway & Workflow Visualizations
Diagram 1: Bioactivity Loss Diagnostic Decision Tree (94 chars)
Diagram 2: Multi-Compound Synergy Enables Activity (92 chars)
The Scientist's Toolkit: Research Reagent Solutions
| Item/Category | Function & Rationale |
|---|---|
| Solid Phase Extraction (SPE) Cartridges (C18, Diol, SCX) | Rapid fractionation of crude extracts for recombination assays; separates compounds by polarity/charge. |
| LC-MS Compatible Buffers (Ammonium Formate/Acetate) | Enable real-time stability profiling and degradation product identification without signal suppression. |
| Stabilizing Additives (Ascorbate, EDTA, BHT) | Protect labile compounds (phenols, terpenes) from oxidative degradation during isolation and storage. |
| Cyclodextrins (HP-β-CD, SBE-β-CD) | Increase aqueous solubility of lipophilic pure compounds for biological testing via inclusion complexation. |
| Inert Atmosphere Vials (Crimp/Septum) | Store purified compounds under argon or nitrogen to prevent oxidation, especially after solvent removal. |
| Analytical & Preparative HPLC Columns | High-resolution separation to isolate single compounds and closely related analogs that may be co-factors. |
| DMSO (Anhydrous, Sterile) | Universal solvent for creating concentrated, sterile stock solutions of pure compounds for cell-based assays. |
Troubleshooting & FAQ Guide
For Researchers Investigating Bioactivity Loss in Compound Isolation
Q1: In our whole-plant extract screens, we observe strong anti-proliferative activity against cancer cell lines. However, upon isolating the three major constituent alkaloids and testing them individually or in a simple reconstituted mixture, we see a >70% loss of efficacy. What are the primary mechanisms for this loss?
A1: The most common culprits are the disruption of synergistic and additive effects. Bioactivity in crude extracts often relies on multi-target network pharmacology, where compounds:
- Synergize: One compound enhances the target binding or cellular uptake of another (pharmacokinetic synergy), or they hit different nodes in a disease-related pathway (pharmacodynamic synergy).
- Additively Interact: Multiple compounds with similar but weak effects sum to produce a strong, significant outcome.
- Provide a Scaffolding Matrix: Minor, non-active constituents in the extract may improve the solubility or stability of the key bioactive compounds, which is lost during purification.
Troubleshooting Step: Perform a systematic fraction re-combination assay. Recombine your isolated compounds in different ratios and combinations, and include a re-constituted "total" fraction. Compare IC50 or growth inhibition values to the original extract.
Q2: Our standard bioassay for anti-inflammatory activity (NO inhibition in macrophages) works perfectly with a fungal culture filtrate but fails after we've fractionated it using our standard HPLC protocol. The activity seems to "vanish." What specific experimental checks should we perform?
A2: This indicates potential compound degradation or missed interactions during fractionation.
Troubleshooting Protocol:
- Immediate Post-Fraction Bioassay: Re-test pooled fractions immediately after drying (under inert gas) and reconstitution. Do not store them for extended periods.
- Check for Chemical Instability:
- pH Sensitivity: Reconstitute fractions and the original extract at different pHs (e.g., 5.0, 7.4, 8.5) and incubate for 1 hour at 37°C before assay.
- Thermal Degradation: Subject a portion of the active fraction to a short heat challenge (e.g., 60°C for 10 mins) and re-test.
- "Spike-In" Experiment: Take your isolated, apparently inactive major compound and "spike" it back into a sub-active concentration of the original crude extract. A return of full activity suggests the crude contains a necessary co-factor.
Q3: We have evidence of synergistic pairs from our recombination studies. How can we definitively prove the mechanism of synergy (e.g., multi-target vs. pharmacokinetic) before investing in complex structural modification?
A3: Implement the following targeted experimental workflows:
Protocol 1: Assessing Pharmacokinetic (PK) Synergy
- Objective: Determine if one compound improves the cellular uptake or stability of another.
- Method: Use LC-MS/MS to quantify intracellular concentrations of each compound when dosed alone vs. in combination over a time course (e.g., 15, 30, 60, 120 mins).
- Key Data: Calculate Area Under the Curve (AUC) for intracellular concentration. A significant increase in AUC for Compound B when co-dosed with Compound A indicates PK-based synergy.
Protocol 2: Mapping Pharmacodynamic (PD) / Multi-Target Synergy
- Objective: Identify if compounds hit different targets in a connected pathway.
- Method:
- Perform target-based assays (e.g., enzyme inhibition, receptor binding) for each isolated compound on suspected pathway targets (e.g., COX-2, 5-LOX, iNOS).
- Use a pathway reporter assay (e.g., NF-κB or AP-1 luciferase reporter cell line). Treat with individual compounds and combinations at sub-effective doses.
- Interpretation: If individual compounds show weak/no activity in the target assays but the combination strongly suppresses the pathway reporter, a multi-target mechanism is likely.
Table 1: Efficacy Loss in Compound Isolation from Plantago ardua Extract
| Sample Type | Assay (IC50, μg/mL) | % Viability (at 50 μg/mL) | Synergy Index (CI)* |
|---|---|---|---|
| Crude Ethanolic Extract | 12.4 ± 1.7 | 22% ± 3 | N/A |
| Isolated Compound A | >100 | 85% ± 5 | N/A |
| Isolated Compound B | 78.5 ± 6.2 | 65% ± 4 | N/A |
| Simple A+B Mixture (1:1) | 45.2 ± 3.8 | 48% ± 4 | 0.92 |
| Reconstituted Full Spectrum | 15.8 ± 2.1 | 25% ± 3 | 0.25 |
CI < 1 indicates synergy; CI ~ 1 indicates additivity; CI > 1 indicates antagonism. Calculated via Chou-Talalay method.
Table 2: Impact of Fractionation Solvents on Bioactivity Recovery
| Fractionation Step | Solvent System | Anti-Biofilm Activity (% Inhibition) | Notes |
|---|---|---|---|
| Crude Broth | N/A | 92% ± 2 | Reference |
| Liquid-Liquid Partition | Ethyl Acetate / H₂O | 88% ± 3 | Good recovery |
| First Normal Phase | Hexane:EtOAc (Gradient) | 15% ± 6 | Major activity loss |
| Second Normal Phase | CH₂Cl₂:MeOH (Gradient) | 5% ± 3 | Activity vanished |
| Activity Rescue | Re-pool of Factions 12-15 & 22-25 | 81% ± 4 | Synergistic pair identified |
Experimental Protocol: Systematic Recombination Bioassay
Objective: To identify synergistic interactions responsible for bioactivity lost during isolation.
Materials:
- Isolated pure compounds (A, B, C, D...)
- Original active crude extract
- Assay plates (96-well)
- Cell line or enzyme target for bioassay
- DMSO (for compound dissolution)
- Multichannel pipettes
Procedure:
- Prepare stock solutions of each isolated compound at a fixed concentration (e.g., 10 mM in DMSO).
- Design Recombination Matrix: Use a checkerboard or fixed-ratio design. For a 4-compound system, test:
- All single compounds.
- All pairwise combinations (A+B, A+C, A+D, B+C...).
- Key triple combinations.
- A full reconstitution (A+B+C+D).
- The crude extract as a control.
- Serially dilute combinations across the assay plate.
- Run your standard bioassay (e.g., cell viability, enzyme inhibition).
- Data Analysis: Calculate Combination Index (CI) using CompuSyn or similar software. CI < 1 indicates synergy.
Visualizations
Diagram 1: Bioactivity Loss in Compound Isolation
Diagram 2: Multi-Target Synergy in a Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Primary Function in Synergy Research |
|---|---|
| Chou-Talalay Software (CompuSyn) | Quantifies drug combination effects (Additivity, Synergy, Antagonism) by calculating the Combination Index (CI). |
| Checkerboard Assay Plates | Enables efficient screening of multiple compound combinations across a matrix of concentrations. |
| LC-MS/MS System | Critical for quantifying intracellular concentrations of compounds to prove pharmacokinetic synergy. |
| Pathway-Specific Reporter Cell Lines (e.g., NF-κB-Luc, AP-1-Luc) | Allows visualization of synergistic inhibition of entire signaling pathways, not just single targets. |
| Isothermal Titration Calorimetry (ITC) | Measures direct binding constants and can identify if one compound alters the binding affinity of another for a target. |
| SPR Biosensor (Surface Plasmon Resonance) | Detects multi-target engagement by a compound mixture on immobilized protein chips. |
| Fraction Library (Pre-plated) | A physical library of all intermediate fractions from isolation, essential for activity tracking and recombination. |
Technical Support Center
Troubleshooting Guides & FAQs
FAQ 1: After isolating my target protein, I observe a >70% loss of catalytic activity compared to crude lysate assays. What are the primary suspects?
- Answer: This is a classic symptom of native environment disruption. The primary suspects are:
- Loss of Essential Cofactors: Metal ions (e.g., Mg²⁺, Zn²⁺), coenzymes (NAD(P)H, ATP), or prosthetic groups may have been stripped during purification.
- Dilution/Removal of Macromolecular Crowders: The intracellular milieu is densely packed (≈300-400 g/L of macromolecules). Isolated proteins in dilute buffers lack this stabilizing "crowding" effect, disrupting folding and complex assembly.
- Disruption of Transient or Weak Protein-Protein/Protein-Lipid Interactions: Many signaling complexes are held together by weak, transient interactions that are lost upon extraction from the membrane or cytosolic matrix.
FAQ 2: How can I systematically identify which missing cofactor is responsible for my enzyme's lost activity?
- Answer: Implement a High-Throughput Cofactor Screen.
- Experimental Protocol:
- Prepare a 96-well plate with your purified, inactive enzyme in a suitable reaction buffer (lacking specific cofactors).
- In separate wells, supplement the reaction mixture with individual candidate cofactors (e.g., MgCl₂, MnCl₂, CaCl₂, ZnSO₄, NAD⁺, NADP⁺, ATP, SAM, PLP) at a range of physiological concentrations (e.g., 0.1 µM to 10 mM).
- Initiate the reaction by adding substrate. Monitor product formation spectrophotometrically or fluorometrically over time.
- Compare initial reaction rates (V₀) to identify which cofactor restores activity. Use the table below to quantify results.
- Experimental Protocol:
Table 1: Example Results from a Cofactor Rescue Screen
| Cofactor Added (1 mM) | Measured V₀ (nmol/min/µg) | Activity (% of Crude Lysate Control) | Interpretation |
|---|---|---|---|
| None (Buffer Only) | 0.5 | 5% | Baseline inactivity |
| MgCl₂ | 8.2 | 82% | Primary Cofactor |
| MnCl₂ | 5.1 | 51% | Partial substitution |
| CaCl₂ | 0.6 | 6% | No effect |
| NAD⁺ | 0.7 | 7% | No effect |
| Crude Lysate Control | 10.0 | 100% | Benchmark |
FAQ 3: My membrane-associated receptor shows no binding affinity for its ligand in isolation. What steps should I take to reconstitute its native function?
- Answer: The issue likely stems from the loss of the lipid membrane environment and associated signaling proteins. Follow this workflow for Functional Membrane Protein Reconstitution.
- Experimental Protocol:
- Solubilize with Native-like Lipids: Use mild detergents (e.g., DDM, digitonin) supplemented with native lipid extracts (e.g., liver polar, brain lipid) during extraction to preserve lipid-protein interactions.
- Purify in Detergent-Lipid Micelles: Include lipids throughout the purification process (e.g., in size-exclusion chromatography buffers).
- Reconstitute into Proteoliposomes: Mix the purified protein with synthetic or native lipid mixtures. Use biobeads or dialysis to remove detergent, forming lipid bilayers containing your receptor.
- Reconstitute Downstream Components: Co-reconstitute purified G-proteins or kinases if studying signal transduction. Assay for ligand binding (e.g., Surface Plasmon Resonance) or downstream activity (e.g., GTPγS binding for GPCRs).
- Experimental Protocol:
Diagram 1: Membrane Protein Activity Loss & Recovery
The Scientist's Toolkit: Research Reagent Solutions for Native Environment Reconstitution
| Reagent / Material | Function & Rationale |
|---|---|
| Native Lipid Extracts (e.g., E. coli Total, Liver Polar) | Provides a physiologically relevant lipid mixture to stabilize membrane proteins and restore lipid-dependent activity during solubilization and reconstitution. |
| Mild Detergents (DDM, Digitonin, LMNG) | Solubilizes membranes while preserving protein-protein complexes and, when used with lipids, can maintain a native-like annular lipid shell. |
| Macromolecular Crowders (Ficoll PM70, Dextran, PEG) | Mimics the high intracellular macromolecule concentration, improving folding stability and promoting weak interaction complex assembly in in vitro assays. |
| Cofactor Cocktails (Cell-Based or Defined) | Pre-mixed solutions of common metal ions and coenzymes to systematically screen for dependencies in purified enzyme systems. |
| Bio-Beads SM-2 | Hydrophobic polystyrene beads used to adsorb detergent, enabling gentle and efficient reconstitution of membrane proteins into lipid bilayers (proteoliposomes). |
| Nanodiscs (MSP Protein / Styrene Maleic Acid Copolymer) | Provides a stable, soluble, and tunable phospholipid bilayer platform to incorporate membrane proteins in a native-like lipid environment for biophysical studies. |
Troubleshooting Guides & FAQs
FAQ 1: My isolated natural compound precipitates upon buffer dilution from DMSO stock. How can I improve solubility without altering bioactivity?
- Answer: Precipitation indicates a solubility shift due to changes in polarity and ionic strength. First, characterize the logP of your compound. For hydrophilic buffers (e.g., PBS), consider:
- Gradual Dilution: Perform serial dilution, adding buffer slowly to the DMSO stock with vigorous vortexing at each step. Do not exceed a final DMSO concentration of 1-5% for cell-based assays.
- Alternative Solvents/Solubilizing Agents: For compounds with logP > 3, use co-solvents like ethanol or propylene glycol (final concentration <1%). For critical assays, use non-ionic surfactants (e.g., 0.01% polysorbate 80) or cyclodextrins (e.g., 2-HP-β-CD at 5-10 mM) to form inclusion complexes.
- Buffer Modification: Slightly increase buffer pH for acidic compounds or decrease pH for basic compounds to promote ionization and aqueous solubility, provided it aligns with your assay's physiological range.
FAQ 2: During bioactivity screening, my compound shows significant activity loss after 24 hours in assay media. Is this a stability or conformation issue?
- Answer: Time-dependent activity loss points to compound instability. Key culprits and diagnostics:
- Hydrolytic Degradation: Check for pH-sensitive functional groups (esters, lactones). Run a simple stability test by HPLC: incubate the compound in assay media at 37°C and sample at 0, 6, 12, 24h. A decreasing parent peak indicates degradation.
- Oxidative Degradation: Suspect this if the compound contains phenols, thiols, or unsaturated bonds. Repeat the stability test under an inert atmosphere (N₂) or add antioxidants like 0.1% ascorbic acid. Compare degradation profiles.
- Conformational Changes (Aggregation): Some compounds form inactive aggregates in aqueous solution. Test by dynamic light scattering (DLS). If aggregates >100 nm are detected, consider adding a low percentage of a benign organic co-solvent or a aggregator-disrupting agent like CHAPS.
FAQ 3: How can I confirm if a conformational change upon solvation is responsible for the loss of target binding?
- Answer: Employ biophysical techniques to probe conformation in the isolation solvent versus the assay buffer.
- Circular Dichroism (CD) Spectroscopy: Compare the CD spectra (far-UV for proteins/peptides; near-UV for small molecule chirality) in both solvents. A shift in peak minima/maxima indicates a conformational change.
- NMR Spectroscopy: ¹H-2D NOESY can identify through-space interactions that differ between solvents, revealing structural rearrangements.
- Differential Scanning Fluorimetry (DSF): If the target protein is available, monitor its melting temperature (Tm) with the compound present in different solvents. A change in ΔTm suggests altered binding due to compound conformation.
Table 1: Common Solubilizing Agents and Their Applications
| Agent | Typical Working Concentration | Mechanism | Best For | Caveat |
|---|---|---|---|---|
| DMSO | 0.1-1% (cell assays) | Universal polar aprotic solvent | Initial stock solutions | Cytotoxic at >1%, can affect membrane permeability |
| 2-Hydroxypropyl-β-Cyclodextrin (HP-β-CD) | 5-20 mM | Forms non-covalent inclusion complexes | Hydrophobic small molecules (LogP >3) | Can weakly extract cholesterol, may require controls |
| Polysorbate 80 (Tween 80) | 0.01-0.1% | Non-ionic surfactant, micelle formation | Moderately lipophilic compounds | Can cause foaming, potential interference in some assays |
| Ethanol | 0.5-2% | Co-solvent, reduces dielectric constant | Compounds stable in alcohols | Evaporation concerns, affects metabolic activity |
| Polyethylene Glycol 400 (PEG 400) | 1-5% | Co-solvent, viscosity enhancer | Improving dissolution kinetics | High viscosity can complicate pipetting |
Table 2: Stability Diagnostic Experiments
| Assay | Protocol Summary | What it Identifies | Key Output Metrics |
|---|---|---|---|
| Forced Degradation (HPLC) | Incubate compound at 37°C in relevant pH buffers (e.g., pH 2, 7.4, 9). Sample at 0, 8, 24, 48h. | Chemical stability (hydrolysis, oxidation) | % Recovery of parent peak; appearance of new degradation peaks. |
| Dynamic Light Scattering (DLS) | Prepare compound at assay concentration in final buffer. Measure particle size distribution. | Physical instability (aggregation, precipitation) | Z-average diameter (d.nm); Polydispersity Index (PDI). |
| Circular Dichroism (CD) | Prepare identical compound concentrations in stock solvent and assay buffer. Scan appropriate UV range. | Conformational shifts (secondary/tertiary structure) | Mean residue ellipticity (MRE) spectra; characteristic peak shifts. |
Experimental Protocols
Protocol 1: Serial Dilution for Solubility Maintenance
- Prepare a 50 mM stock of your compound in 100% DMSO.
- Prepare your aqueous assay buffer (e.g., PBS, pH 7.4).
- In a microcentrifuge tube, create a 1:10 intermediate dilution: Add 10 µL of 50 mM stock to 90 µL of pure DMSO. Vortex for 10 seconds.
- For the working solution: Add 10 µL of the intermediate dilution slowly (dropwise over 10 seconds) to 990 µL of pre-warmed assay buffer while vortexing continuously. Final concentrations: 50 µM compound, 1% DMSO.
- Visually inspect for precipitation. If clear, use immediately.
Protocol 2: Differential Scanning Fluorimetry (DSF) for Binding Assessment
- Sample Prep: Prepare 20 µL samples in a 96-well PCR plate containing: 5 µM target protein, 5X SYPRO Orange dye, compound (at 10x desired final concentration in matched solvent), and assay buffer. Include a solvent-only control.
- Run: Seal plate, centrifuge briefly. Use a real-time PCR machine with a FRET filter set. Ramp temperature from 25°C to 95°C at a rate of 1°C/min.
- Analysis: Plot fluorescence (RFU) vs. Temperature. Determine the melting temperature (Tm) as the inflection point of the sigmoidal curve for each condition. A shift in Tm (ΔTm) > 1°C between compound and control suggests binding/stabilization.
Visualizations
Title: Physicochemical Shifts Leading to Bioactivity Loss
Title: Diagnostic & Mitigation Workflow for Activity Loss
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Primary Function | Critical Application Note |
|---|---|---|
| 2-HP-β-Cyclodextrin | Molecular encapsulant to enhance aqueous solubility of hydrophobic compounds. | Use in binding assays where DMSO is problematic. Always run a vehicle control with identical CD concentration. |
| SYPRO Orange Dye | Environment-sensitive fluorescent dye for protein denaturation detection in DSF. | Optimize dye dilution (typically 5-10X) to avoid signal quenching. Protect from light. |
| Deuterated Solvents (DMSO-d6, D2O) | NMR-compatible solvents for comparing compound conformation in different environments. | For direct comparison, acquire NMR spectra in pure deuterated organic solvent vs. buffer/D2O mixtures. |
| Polysorbate 80 (Tween 80) | Non-ionic surfactant to prevent compound adsorption and micelle-mediated solubilization. | Filter-sterilize (0.22 µm) surfactant stock solutions. Be aware of potential effects on membrane proteins. |
| CHAPS Detergent | Zwitterionic detergent used to disrupt non-covalent, inactive compound aggregates. | Use at low concentrations (e.g., 0.1%) in activity assays to test for aggregate-related false negatives. |
| HPLC vials with pre-slit septa | Sample integrity for stability-indicating chromatographic analysis. | Essential for preventing evaporation and ensuring accurate quantification of parent compound over time. |
Troubleshooting Guide & FAQs
General Compound Isolation & Bioactivity
Q1: My isolated natural product shows significantly lower antimicrobial activity in the pure state compared to the crude extract. What are the primary causes? A: This is a common issue in advancing bioactivity lost during isolation. Key causes include:
- Loss of Synergistic Partners: The pure compound may have acted synergistically with other components in the crude extract. Its isolated effect is inherently weaker.
- Compound Instability: The isolation process (pH changes, solvent exposure, chromatography) can degrade the active compound or modify its functional groups.
- Altered Solubility/Pharmacokinetics: The compound's formulation in the crude extract (e.g., in a lipid milieu) may have provided better bioavailability than your test solution (e.g., DMSO/aqueous buffer).
Q2: During anticancer screening, my compound is active in cell culture but fails in vivo. What should I troubleshoot first? A: Focus on ADMET (Absorption, Distribution, Metabolism, Excretion, Toxicity) properties.
- Pharmacokinetics: Check plasma stability and half-life. The compound may be rapidly metabolized or cleared in vivo.
- Solubility & Formulation: The in vitro DMSO stock may not translate to a soluble, bioavailable form in vivo. Reformulate using acceptable vehicles (e.g., PEG, cyclodextrins).
- Metabolic Activation: Some compounds require liver metabolism to become active. Test activity in the presence of microsomal fractions.
Q3: I suspect my antimicrobial compound's activity is due to synergy, not a single agent. How can I design an experiment to confirm this? A: Perform a Checkerboard Assay to calculate the Fractional Inhibitory Concentration Index (FICI).
| FICI Value | Interpretation |
|---|---|
| ≤ 0.5 | Synergy |
| >0.5 - 4 | Additivity / No Interaction |
| > 4 | Antagonism |
- Protocol: In a 96-well plate, titrate Compound A along the rows and Compound B (or crude fraction) along the columns. Inoculate with a standardized microbial inoculum. After incubation, measure growth (OD600). Calculate FICI = (MIC of A in combo / MIC of A alone) + (MIC of B in combo / MIC of B alone).
Technical & Experimental Issues
Q4: My compound precipitates out of solution in cell culture media, confounding my bioassay results. How can I address this? A:
- Use a Compatible Solvent: Ensure the final concentration of DMSO does not exceed 0.1-1% (v/v). For water-insoluble compounds, consider alternative solvents like ethanol or use solvent-free methods.
- Employ a Solubilizing Agent: Use bovine serum albumin (BSA, 0.1-1%) or cyclodextrins (e.g., HP-β-CD) to enhance aqueous solubility. Include a vehicle control with the same concentration of solubilizer.
- Sonication & Warming: Briefly sonicate and warm the stock solution in a water bath (37°C) before diluting into pre-warmed media.
Q5: How can I quickly assess if the loss of anticancer activity is due to apoptosis induction failure? A: Perform a Caspase-3/7 Activation Assay as an early apoptotic marker.
- Protocol: Seed cancer cells in a white-walled 96-well plate. Treat with your compound for 6-24 hours. Add a luminescent caspase-3/7 substrate (e.g., Caspase-Glo Reagent). Incubate for 30-60 minutes and measure luminescence. Compare to a known apoptosis inducer (e.g., staurosporine) as a positive control.
Experimental Protocols
Protocol 1: Checkerboard Synergy Assay for Antimicrobials
Objective: Determine synergistic interactions between a purified compound and a crude fraction or another antibiotic.
- Prepare Mueller-Hinton Broth (MHB) for bacteria or RPMI-1640 for fungi.
- Standardize the microbial inoculum to 0.5 McFarland, then dilute to ~5 x 10^5 CFU/mL in broth.
- In a sterile 96-well plate, add 50 µL of broth to all wells.
- Perform serial 2-fold dilutions of Compound A down the rows (e.g., columns 1-12). Add 50 µL per well.
- Perform serial 2-fold dilutions of Compound B (or fraction) across the columns (e.g., rows A-H). Add 50 µL per well. This creates all combinations.
- Add 50 µL of the standardized inoculum to each well. Final volume = 150 µL. Columns 11-12 are growth controls (no compound). Column 12 is a sterility control (no inoculum).
- Incubate statically at 37°C for 18-24 hours.
- Read MIC as the lowest concentration with no visible growth. Calculate FICI.
Protocol 2: ATP-Based Viability Assay (CellTiter-Glo) for Anticancer Screening
Objective: Quantify viable cells after compound treatment based on cellular ATP levels.
- Seed cancer cells in a 96-well tissue culture plate at an optimal density (e.g., 2000-5000 cells/well in 90 µL medium). Incubate overnight.
- Prepare compound dilutions in complete medium. Add 10 µL to respective wells for a 10X final concentration. Include vehicle and positive control (e.g., 10 µM Staurosporine).
- Incubate for desired time (e.g., 48-72 hours).
- Equilibrate CellTiter-Glo reagent and plate to room temperature (20-25°C) for 30 minutes.
- Add 100 µL of reagent directly to each well.
- Mix on an orbital shaker for 2 minutes to induce cell lysis.
- Allow the plate to incubate at RT for 10 minutes to stabilize luminescent signal.
- Record luminescence using a plate reader. Plot % viability vs. log[compound].
Visualizations
Flowchart: Bioactivity Loss During Compound Isolation
Pathway: Intrinsic Apoptosis Signaling for Compound Validation
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Primary Function in This Context |
|---|---|
| HP-β-Cyclodextrin | A solubilizing agent used to enhance the aqueous solubility of hydrophobic compounds in bioassays, preventing precipitation and false negatives. |
| Caspase-Glo 3/7 Assay | A luminescent, homogeneous assay to measure caspase-3 and -7 activity as a key marker of apoptosis induction in treated cells. |
| CellTiter-Glo Luminescent Assay | A gold-standard, ATP-based method for quantifying the number of viable cells in culture post-treatment with test compounds. |
| S9 Liver Microsomal Fraction | Used in metabolic stability assays to predict Phase I hepatic metabolism and identify if a compound requires metabolic activation. |
| 96-Well Checkerboard Plate | A specialized microplate layout facilitating the systematic testing of all concentration combinations of two agents for synergy studies. |
| Matrigel Basement Membrane Matrix | Used in in vitro invasion assays and for creating more physiologically relevant 3D cell culture models for anticancer testing. |
| Resazurin Sodium Salt | A redox indicator used in alamarBlue assays for measuring cell viability and proliferation in both antimicrobial and anticancer screens. |
Recovery Protocols: Methodologies to Restore and Enhance Bioactivity
Frequently Asked Questions (FAQs)
Q1: I consistently observe a significant drop in total bioactivity when moving from a bioactive crude extract to isolated pure compounds. What are the primary causes and how can I mitigate this? A1: This "lost bioactivity" is a central challenge. Primary causes and mitigations are summarized in the table below.
| Cause of Bioactivity Loss | Mechanism | Mitigation Strategy |
|---|---|---|
| Synergistic Effects Lost | Bioactivity relies on multiple compounds acting together. Isolation removes complementary components. | Employ systematic combination studies. Recombine fractions and test for restored activity. |
| Compound Instability | The pure compound may degrade during isolation (pH changes, light, oxidation) or lose activity outside its native matrix. | Optimize isolation buffers (use antioxidants, chelators). Minimize processing steps and time. |
| Non-Specific Binding/Matrix Effect | Activity in crude extract may depend on non-specific protein binding or co-factors present in the mixture. | Replicate assay conditions with added inert protein (e.g., BSA) or synthetic lipid vesicles. |
| Incorrect Bioassay | The assay may measure a complex phenotypic response not linked to a single target, misguiding isolation. | Use orthogonal bioassays (cell-based + target-based) to guide fractionation. |
| Pharmacokinetic Effects | The crude extract may contain compounds that improve bioavailability (e.g., solubility, membrane penetration) of the active. | Include parallel ADMET screening (solubility, permeability) early in the isolation workflow. |
Q2: During bioactivity-guided fractionation, my active fraction becomes inactive after the next chromatographic step, even though HPLC shows a pure compound. What went wrong? A2: This indicates a critical point of loss. Follow this troubleshooting protocol:
- Immediate Re-analysis: Re-test the "inactive" pure compound and the immediately preceding active fraction side-by-side in your bioassay to confirm the result.
- Stability Check: Perform a time-course stability test. Incubate the pure compound in the elution solvent and assay buffer, then measure activity at 0, 2, 4, 8, 24 hours.
- Recombination Test: This is crucial. Spiking the pure compound back into an inactive matrix (or a fraction from earlier in the process) can reveal if a synergist was lost.
- Structural Confirmation: Use LC-MS/MS to confirm the isolated compound did not undergo structural modification (e.g., oxidation, hydrolysis) on-column.
Q3: What are the best practices for selecting and validating a bioassay to guide fractionation to avoid misleading results? A3: A robust bioassay is foundational. See the validation protocol below.
Protocol: Bioassay Validation for Fractionation Guidance
- Define Criteria: Select an assay relevant to the therapeutic hypothesis (e.g., anti-inflammatory = COX-2 inhibition + cytokine reduction in cells).
- Z'-Factor Test: Determine the assay's robustness using a positive control (inhibitor) and negative control (vehicle). A Z' factor > 0.5 is essential for reliable hit identification.
- Dose-Response Validation: Ensure your crude extract produces a clear, reproducible dose-response curve (IC50/EC50).
- Interference Testing: Test common extraction solvents (DMSO, MeOH, EtOAc) and fractionation buffers in the assay at their maximum expected concentration to rule out artifactual inhibition/activation.
- Orthogonal Assay: Establish a secondary, mechanistically different assay to confirm hits from the primary screen.
Q4: Can you provide a standard workflow that integrates strategies to minimize bioactivity loss? A4: Yes. The following diagram outlines an integrated workflow.
Integrated Workflow to Minimize Bioactivity Loss
Q5: How do I design experiments to test for lost synergistic interactions? A5: Follow this systematic combination protocol.
Protocol: Testing for Synergistic Interactions Post-Isolation
- Prepare Samples: Isolated pure compound (P), inactive or marginally active fractions (F1, F2...Fn) from preceding separation steps.
- Design Matrix: Create a combination matrix. Test P alone, each F alone, and P + each F at a fixed ratio (e.g., based on original crude extract weight).
- Bioassay: Test all samples in your primary bioassay.
- Data Analysis: Calculate Combination Index (CI) using the Chou-Talalay method. CI < 1 indicates synergy, CI = 1 additivity, CI > 1 antagonism.
- Identification: For synergistic pairs (P+Fx), subject Fx to further analysis (metabolomics) to identify the synergistic partner(s).
The Scientist's Toolkit: Key Reagent Solutions
| Reagent / Material | Primary Function in This Context |
|---|---|
| Solid Phase Extraction (SPE) Cartridges (C18, Diol, SCX) | Rapid desalting and pre-fractionation of crude extracts with minimal solvent use, reducing degradation time. |
| Sephadex LH-20 | Size-exclusion and adsorption chromatography for gentle separation of natural products based on molecular size/polarity. |
| Prefilled Silica/C18 Flash Columns | For medium-pressure liquid chromatography (MPLC) to scale up separation of active fractions reliably. |
| LC-MS Grade Solvents & Modifiers | High-purity solvents (MeCN, MeOH, H₂O) and modifiers (TFA, Formic Acid, NH₄OAc) for HPLC to prevent artifact formation. |
| Deuterated NMR Solvents (e.g., DMSO-d6, CD3OD) | For structural elucidation of unstable compounds; some offer stabilizing effects. |
| Stabilizer Cocktails | Ready-to-use mixes of antioxidants (e.g., BHT, ascorbic acid), chelators (EDTA), and protease inhibitors to add to extraction/isolation buffers. |
| 96-Well Assay Plates (Cell-culture treated) | For high-throughput bioactivity screening of numerous fractions with minimal sample consumption. |
| Bioactive Compound Standards (e.g., COX-2 Inhibitor, Staurosporine) | Essential positive controls for bioassay validation and monitoring performance during long fractionation projects. |
Pathway Diagram: Common Bioactivity Loss During Isolation
Mechanisms of Bioactivity Loss During Compound Isolation
Quantitative Data Summary: Common Pitfalls
| Experimental Stage | Typical Activity Loss (Reported Range) | Major Contributing Factor |
|---|---|---|
| Crude Extract to First Fraction | 10-40% | Poor fractionation resolution leading to split of synergistic pairs. |
| Between Chromatographic Steps | 20-60% | Compound degradation due to pH, adsorption to stationary phase, or oxidation. |
| Final Pure Compound vs. Original Extract | 50-100% | Loss of synergism is the most cited factor, accounting for >70% of major losses in antimicrobial and anticancer studies. |
| Mitigation Impact | Activity Recovery Range | Strategy Employed |
| Recombination Studies | 30-95% | Re-introducing an "inactive" fraction to the pure compound. |
| Use of Stabilizers | 15-50% | Adding antioxidants & chelators to all solvents/buffers. |
| Accelerated Isolation | 10-30% | Reducing total processing time from weeks to days. |
Technical Support Center: Troubleshooting & FAQs
FAQ 1: Why is the bioactivity of my reconstituted mixture significantly lower than the original crude extract?
Answer: This is a common issue. Key factors include:
- Incorrect Ratio: The original synergistic ratio of isolated compounds may not have been accurately determined. Use systematic combinatorial screening (e.g., fractional factorial design) to re-optimize.
- Missing Minor Components: Bioactivity often depends on trace compounds not identified during isolation. Consider LC-MS or NMR-based metabolomics to profile the original extract and compare it to your reconstituted mix.
- Compound Degradation: Isolated pure compounds may degrade over time. Check compound stability under storage conditions and use fresh stock solutions.
FAQ 2: How do I determine the optimal ratios for reconstituting isolated compounds?
Answer: Employ Design of Experiments (DoE) methodologies. A central composite design is highly effective for modeling synergistic interactions. Prepare stock solutions of each isolated compound and use the DoE matrix to create mixtures across a concentration matrix. Assay for bioactivity and use response surface modeling to identify the optimal interaction landscape.
FAQ 3: My reconstituted mixture shows antagonism instead of synergy. What could be the cause?
Answer: Antagonism can arise from:
- Non-physiological Ratios: The concentrations used may be outside the therapeutic window, leading to inhibitory cross-talk.
- Solvent/Vehicle Incompatibility: Different solubility of isolated compounds may require use of solvents that affect biological target integrity. Ensure all compounds are in a biocompatible vehicle (e.g., DMSO < 0.1% final concentration).
- Oxidation or Interaction in Solution: Compounds may react with each other in the mixture. Analyze the mixture stability over time using HPLC.
Key Experimental Protocol: Systematic Reconstitution & Synergy Screening
Objective: To quantitatively rebuild and optimize a bioactive mixture from isolated compounds and calculate synergy scores.
Materials:
- Purified compounds (A, B, C...N) from the original bioactive extract.
- Cell-based or enzymatic bioassay system relevant to the original activity.
- Liquid handling robot or multichannel pipettes for high-throughput mixing.
- 384-well assay plates.
- Software for DoE (e.g., JMP, Design-Expert) and synergy calculation (e.g., Combenefit, SynergyFinder).
Methodology:
- Stock Solution Preparation: Prepare concentrated stock solutions of each compound in a compatible solvent. Determine maximum non-toxic concentration for each.
- Experimental Design: Generate a mixture matrix using a checkerboard or fixed-ratio design. For 3 compounds, a typical matrix includes serial dilutions of each compound alone and in all possible combinations.
- Plate Setup: Using automation, dispense compounds and combinations into assay plates according to the design matrix. Include controls (vehicle, positive inhibitor).
- Bioassay: Add assay reagents (cells, enzyme substrate) and incubate under standard conditions. Measure endpoint (e.g., luminescence, fluorescence, absorbance).
- Data Analysis:
- Calculate % inhibition/activation for each well.
- Model expected additive effect using the Bliss Independence or Loewe Additivity model.
- Calculate synergy scores (ΔBliss or Combination Index) for each mixture combination.
- Visualize results in an interaction landscape plot.
Research Reagent Solutions
| Reagent / Material | Function in Reconstitution Experiments |
|---|---|
| Dimethyl Sulfoxide (DMSO), HPLC Grade | Universal solvent for preparing concentrated, stable stock solutions of diverse organic compounds. |
| CellTiter-Glo Luminescent Assay | Robust, homogeneous cell viability assay to measure cytotoxicity and proliferative bioactivity of mixtures. |
| Phosphatidylcholine Liposomes | Membrane models to assess the impact of lipid partitioning on compound interaction and delivery. |
| HP-β-Cyclodextrin | Solubility enhancer for poorly water-soluble compounds, ensuring they remain in solution during biological testing. |
| LC-MS/MS System with PDA | Critical for chemical profiling of original extract and quality control of reconstituted mixtures to verify composition. |
| SynergyFinder Web Tool | Open-source software for analyzing drug combination data and calculating multiple synergy scores (Bliss, Loewe, HSA). |
Table 1: Example Synergy Scores (ΔBliss) for Ternary Mixture
| Compound A (µM) | Compound B (µM) | Compound C (µM) | Observed Inhibition (%) | Expected Additive Inhibition (%) | ΔBliss Score |
|---|---|---|---|---|---|
| 1.0 | 0 | 0 | 15 | 15 | 0 |
| 0 | 5.0 | 0 | 20 | 20 | 0 |
| 0 | 0 | 10.0 | 10 | 10 | 0 |
| 1.0 | 5.0 | 0 | 50 | 32 | +18 |
| 1.0 | 0 | 10.0 | 40 | 23.5 | +16.5 |
| 0 | 5.0 | 10.0 | 45 | 28 | +17 |
| 1.0 | 5.0 | 10.0 | 85 | 46 | +39 |
A positive ΔBliss score indicates synergy. The ternary mixture shows strong synergistic bioactivity.
Table 2: Troubleshooting Common Experimental Failures
| Symptom | Possible Cause | Verification Test | Solution |
|---|---|---|---|
| No activity in any mixture | Compound stocks degraded | Re-analyze stocks via HPLC vs. standard | Prepare fresh stocks; use stabilizers; store at -80°C. |
| High background noise in assay | Solvent (DMSO) concentration too high | Run vehicle control gradient | Ensure final DMSO ≤0.1%. Use alternative solubilizers. |
| Inconsistent replicate data | Manual pipetting errors in mixture prep | Use dye dilution test for pipette accuracy | Implement liquid handler for mixture preparation. |
| Activity plateau at low level | Missing a critical co-factor from extract | Add back fractions of inactive extract | Use bioassay-guided fractionation to find missing element. |
Visualizations
Title: Workflow for Reconstituting Bioactive Synergistic Mixtures
Title: Synergistic Pro-Apoptotic Pathway of a Ternary Mixture
Technical Support Center
Troubleshooting Guides & FAQs
Liposome Encapsulation
Q1: My liposomal formulation has very low encapsulation efficiency (EE%) for my hydrophobic bioactive compound, resulting in a significant loss of bioactivity. What could be the issue?
- A: Low EE% for hydrophobic compounds is often due to improper lipid selection or preparation method. The compound may be partitioning into the bilayer but leaking during downstream processing.
- Troubleshooting Steps:
- Increase Lipid-to-Drug Ratio: Try a higher molar ratio (e.g., from 10:1 to 50:1) to provide more bilayer capacity.
- Optimize Lipid Composition: Incorporate high-phase-transition-temperature lipids (e.g., DSPC) and cholesterol (up to 45 mol%) to improve bilayer rigidity and reduce leakage.
- Change Preparation Method: Switch from thin-film hydration to an active loading method (e.g., pH gradient or ammonium sulfate) if applicable, or use solvent injection techniques for better hydrophobic incorporation.
- Purification Check: Ensure your separation method (e.g., size exclusion chromatography) is not disrupting the liposomes. Consider using gentler methods like dialysis.
Q2: My liposome suspension shows aggregation or fusion upon storage. How can I improve physical stability?
- A: Aggregation indicates instability in the formulation, which can lead to inconsistent dosing and bioactivity.
- Troubleshooting Steps:
- Surface Charge: Incorporate a charged lipid (e.g., 5-10 mol% DOTAP for positive, DOPG for negative charge) to create electrostatic repulsion between vesicles.
- Steric Stabilization: Add 5-10 mol% of a PEGylated lipid (e.g., DSPE-PEG2000) to create a hydrophilic steric barrier.
- Storage Conditions: Store in isotonic buffer (e.g., sucrose or HEPES-buffered saline) at 4°C. Avoid freezing without cryoprotectants (e.g., trehalose).
- Size Control: Ensure liposomes are homogenized (e.g., via extrusion through 100 nm or 200 nm membranes) to a uniform size, which improves stability.
Polymeric Nanoparticles
Q3: My PLGA nanoparticle formulation has a high burst release in vitro, not the sustained release profile needed to mimic prolonged bioactivity.
- A: A high initial burst release is typically caused by drug adsorbed on or near the nanoparticle surface.
- Troubleshooting Steps:
- Optimize Emulsion Process: Increase the homogenization speed/time to form smaller primary emulsion droplets, leading to more homogeneous drug distribution.
- Adjust Organic Phase: Use a less water-miscible organic solvent (e.g., dichloromethane instead of ethyl acetate) to slow the diffusion process.
- Modify PLGA Properties: Use a higher molecular weight PLGA (e.g., 75-100 kDa) or a PLGA with a higher lactide:glycolide ratio (e.g., 75:25) for slower degradation.
- Add a Coating: Apply a stabilizing coating (e.g., poloxamer, chitosan) via post-formulation incubation to create a diffusion barrier.
Q4: My nanoparticle yield after synthesis and purification is very low, making it difficult to recover enough of my isolated compound for testing.
- A: Low yield is often a result of losses during purification or inefficient particle recovery.
- Troubleshooting Steps:
- Centrifugation Optimization: For centrifugation-based washing, optimize the g-force and time. Too low will not pellet all particles; too high may make the pellet too compact and difficult to re-disperse.
- Switch Purification Method: Consider using tangential flow filtration (TFF) for higher recovery yields, especially for smaller nanoparticles (<100 nm).
- Concentration Step: After purification, use a centrifugal concentrator (with an appropriate MWCO) to concentrate the sample without pelleting.
- Check for Aggregation: Aggregated particles may be lost during filtration steps. Re-optimize formulation for colloidal stability (see Q2).
Cyclodextrin Complexation
Q5: My phase-solubility diagram for the cyclodextrin (CD)-compound complex shows an A~L~-type curve, suggesting limited complexation and poor solubility enhancement.
- A: An A~L~-type (linear) curve indicates a 1:1 stoichiometry but with a low stability constant (K~1:1~), meaning the complex is weak.
- Troubleshooting Steps:
- CD Selection: Switch to a derivatized CD with more suitable cavity size or substituents. For hydrophobic compounds, try Sulfobutylether-β-CD (SBE-β-CD) or randomly methylated-β-CD (RM-β-CD).
- Environmental Conditions: Perform complexation at a controlled temperature (e.g., 25°C) and adjust the pH to ensure your compound is in its unionized form if that favors complexation.
- Preparation Method: Use a more rigorous method like co-evaporation or kneading instead of simple physical mixing to improve complex formation.
- Consider Ternary Complexes: Add a small, complementary third component (e.g., a water-soluble polymer like PVP or a carboxylic acid) that can enhance complex stability.
Q6: My bioactive compound precipitates out of solution upon dilution of the cyclodextrin complex during in vitro assays, confounding bioactivity results.
- A: Precipitation upon dilution occurs when the complex dissociates and the free drug concentration exceeds its solubility in the new medium.
- Troubleshooting Steps:
- Increase CD Concentration: If assay conditions allow, maintain a low concentration of free CD in the final assay medium to shift the equilibrium and keep the drug in solution.
- Pre-equilibrate Assay Media: Prepare your cell culture or assay media with the same concentration of CD as in your stock complex solution before dilution.
- Formulate as a Solid Dosage: If for in vivo use, lyophilize the complex and reconstitute it directly into the full volume of the final administration vehicle.
Quantitative Data Summary
Table 1: Comparative Overview of Drug Delivery Systems for Bioactivity Recovery
| Parameter | Liposomes | Polymeric Nanoparticles (PLGA) | Cyclodextrin Complexes |
|---|---|---|---|
| Typical Size Range | 50 nm - 5 μm | 50 nm - 500 nm | 1 - 2 nm (Molecular Complex) |
| Encapsulation Efficiency | Moderate-High (for suited drugs) | Moderate-High | Very High (for formable complexes) |
| Drug Loading Capacity | Low-Moderate (1-10%) | Moderate (1-30%) | Low (5-20% w/w) |
| Release Profile | Biphasic (burst + sustained) | Triphasic (burst + degradation-controlled) | Instantaneous upon dissociation |
| Key Stability Challenge | Oxidation, hydrolysis, aggregation | Hydrolytic degradation, aggregation | Precipitation upon dilution |
| Scalability | Moderate (GMP possible) | High (well-established) | Very High |
Table 2: Common Experimental Characterization Methods & Target Values
| Characterization | Method | Target/Indicator of Success | ||
|---|---|---|---|---|
| Size & PDI | Dynamic Light Scattering (DLS) | Liposomes/NPs: 80-200 nm, PDI < 0.2. Stable over time. | ||
| Surface Charge | Zeta Potential (ζ) | ζ | > 20 mV for good electrostatic stability. | |
| Encapsulation Efficiency | Centrifugation/Filter separation, HPLC | > 70% for hydrophobic compounds. | ||
| Complexation Efficiency | Phase-Solubility Study | A~P~-type curve with high K~1:1~ stability constant. | ||
| In Vitro Release | Dialysis in sink conditions, HPLC | Matches desired profile (e.g., <30% burst in 24h). |
Detailed Experimental Protocols
Protocol 1: Thin-Film Hydration for Liposome Preparation
- Dissolve: Dissolve lipids (e.g., DPPC:Cholesterol:DSPE-PEG2000 at 55:40:5 molar ratio) and hydrophobic drug in chloroform in a round-bottom flask.
- Form Film: Rotate flask under reduced pressure (using rotary evaporator, 40°C) to form a thin, dry lipid film.
- Hydrate: Hydrate film with aqueous buffer (e.g., PBS, pH 7.4) at temperature above lipid phase transition (e.g., 55°C for DPPC) with vigorous agitation for 1 hour.
- Size Reduction: Subject the multilamellar vesicle suspension to 10 freeze-thaw cycles (liquid N2/55°C water bath). Then extrude 21 times through polycarbonate membranes (e.g., 100 nm pore size) using a mini-extruder.
- Purify: Separate free drug from liposomes using size exclusion chromatography (Sephadex G-50) or dialysis.
Protocol 2: Single Emulsion Solvent Evaporation for PLGA Nanoparticles
- Oil Phase: Dissolve 50 mg PLGA (e.g., 50:50, 24-38 kDa) and 5 mg drug in 2 mL dichloromethane (DCM).
- Emulsify: Add the organic phase to 8 mL of 1-5% (w/v) poly(vinyl alcohol) (PVA) aqueous solution. Homogenize (e.g., 10,000 rpm, 2 min) to form an oil-in-water (O/W) emulsion.
- Evaporate: Stir the emulsion magnetically at room temperature overnight to evaporate DCM and harden nanoparticles.
- Collect & Wash: Centrifuge the suspension at 20,000 x g for 20 min. Discard supernatant, re-suspend pellet in distilled water, and repeat wash twice to remove PVA.
- Lyophilize: Re-suspend final pellet in a small volume of water, add cryoprotectant (e.g., 5% trehalose), freeze at -80°C, and lyophilize for 48h for storage.
Mandatory Visualizations
Diagram 1: Thesis Context - Recovering Bioactivity with Delivery Systems
Diagram 2: Experimental Workflow with Feedback Loop
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Formulation & Characterization
| Reagent/Material | Function & Application |
|---|---|
| DSPC (1,2-distearoyl-sn-glycero-3-phosphocholine) | High Tm lipid for forming rigid, stable liposome bilayers. Reduces drug leakage. |
| PLGA (50:50, 24-38 kDa) | Biodegradable polymer for nanoparticles. Provides sustained, degradation-controlled release. |
| Sulfobutylether-β-Cyclodextrin (SBE-β-CD) | Anionic, water-soluble CD derivative. Enhances solubility & stability of cationic/hydrophobic drugs. |
| Cholesterol (Pharma Grade) | Incorporated into liposomes (up to 45 mol%) to improve membrane stability and reduce permeability. |
| DSPE-PEG2000 | PEGylated lipid for conferring "stealth" properties and prolonging circulation time of liposomes/nanoparticles. |
| Polyvinyl Alcohol (PVA, 87-89% hydrolyzed) | Common stabilizer & emulsifier in PLGA NP preparation. Controls particle size and prevents aggregation. |
| Trehalose Dihydrate | Cryoprotectant for lyophilization of liposomes and nanoparticles. Preserves size and stability upon reconstitution. |
| Dialysis Tubing (e.g., 10 kDa MWCO) | Purifies nanoparticles and liposomes from unencapsulated drug and free small molecules. |
| Polycarbonate Membrane Extruder (100 nm) | Produces uniform, monodisperse liposomes and some nanoparticles via size extrusion. |
Adjuvant and Cofactor Supplementation Strategies
Technical Support Center
Frequently Asked Questions & Troubleshooting
Q1: I have isolated a natural compound that showed promising activity in a crude extract, but the purified compound is inactive in my cell-based assay. What are my first steps? A: This is a classic sign of lost bioactivity due to cofactor depletion or disrupted synergism. First, verify the integrity of your purified compound via HPLC and mass spectrometry to rule out degradation. If intact, proceed to a systematic adjuvant screen. Begin by supplementing the assay medium with a cocktail of common enzyme cofactors (e.g., NAD+, NADP+, Mg²⁺, ATP at 1-10 µM) and essential metal ions (e.g., Zn²⁺, Fe²⁺, Cu²⁺ at physiologically relevant, non-toxic concentrations). Run a pilot dose-response of your compound with and without this baseline cocktail.
Q2: How do I distinguish between a true pharmacological synergist (adjuvant) and a simple, non-specific activity enhancer? A: Conduct a matrixed combination experiment. Titrate both your primary compound and the putative adjuvant independently and in combination. Calculate the Combination Index (CI) using the Chou-Talalay method. A CI < 1 indicates synergism. Additionally, test the adjuvant alone at all concentrations used; it should exhibit minimal to no intrinsic activity at those doses. True adjuvants often show a threshold effect.
Q3: My compound requires a redox-active cofactor (e.g., FAD, CoQ10) that is unstable in culture medium. How can I ensure consistent delivery? A: Instability is common. Consider these solutions:
- Use stabilized analogs: Utilize more stable, cell-permeable prodrug forms (e.g., ubiquinol for CoQ10).
- Pre-condition cells: Pre-incubate cells with the cofactor for 24h prior to compound addition to allow cellular uptake and integration.
- Employ encapsulation: Use liposomal or cyclodextrin-based delivery systems to protect the cofactor. Include a negative control with empty vesicles.
- Continuous infusion: For critical, short-lived cofactors, use a perfusion system or medium exchange protocol.
Q4: In an in vivo model, how can I determine the optimal dosing schedule for a compound-adjuvant pair? A: Pharmacokinetic (PK) profiling is essential. First, establish the individual PK curves for both agents. The adjuvant should be dosed to ensure its peak concentration or area under the curve overlaps with the therapeutic window of the primary compound. Often, this requires administering the adjuvant slightly before or concurrently with the primary drug. A staggered dosing study is critical.
Experimental Protocol: Systematic Adjuvant Rescue Screening
Objective: To identify exogenous cofactors or adjuvants that restore the bioactivity of a purified, inactive compound in a cell-based phenotypic assay.
Materials:
- Purified test compound (inactive)
- Cell line relevant to the original crude extract's bioactivity
- Assay-ready plates (e.g., 96-well)
- Library of potential adjuvants/cofactors (see Reagent Solutions table)
- DMSO (vehicle control)
- Cell viability/activity assay kit (e.g., ATP-based, reporter gene)
Methodology:
- Plate Cells: Seed cells at optimal density in complete growth medium and allow to adhere for 24h.
- Prepare Adjuvant Plates: Using a liquid handler, pre-dispense a matrix of individual adjuvants (at 2x final concentration in serum-free medium) into plates. Include vehicle-only and positive control (crude extract, if available) wells.
- Compound Addition: Prepare a 2x solution of your purified test compound in serum-free medium. Add an equal volume to the adjuvant plate, resulting in 1x final concentrations of both. Final DMSO concentration must be consistent (<0.5%).
- Treatment: Aspirate growth medium from cell plate and transfer the compound-adjuvant mix from the preparation plate.
- Incubate: Incubate for the desired assay duration (e.g., 48h).
- Assay: Perform the endpoint assay (e.g., luminescence for viability).
- Analysis: Normalize data to vehicle control. Identify adjuvants causing a statistically significant (p<0.01) increase in activity compared to the compound alone.
Data Presentation: Adjuvant Screening Results for Compound X in HepG2 Cells
Table 1: Effect of Cofactor Supplementation on the Bioactivity of Purified Compound X (10 µM). Activity measured as % cell viability inhibition relative to crude extract control (mean ± SD, n=6).
| Supplement Class | Specific Adjuvant | Concentration | Activity with Compound X | Adjuvant Alone Activity | p-value vs. Compound X alone |
|---|---|---|---|---|---|
| None (Control) | Vehicle (DMSO) | 0.1% | 5.2% ± 1.8% | 1.1% ± 0.9% | -- |
| Electron Carrier | NADH | 10 µM | 7.5% ± 2.1% | 0.8% ± 1.1% | 0.12 |
| Electron Carrier | FAD | 5 µM | 48.3% ± 5.6% | 2.3% ± 1.4% | <0.001 |
| Metal Ion | MgCl₂ | 100 µM | 8.9% ± 2.4% | 0.5% ± 0.7% | 0.08 |
| Metal Ion | ZnSO₄ | 10 µM | 15.2% ± 3.1% | 3.1% ± 1.8% | 0.02 |
| Antioxidant | Reduced Glutathione | 50 µM | 22.4% ± 4.2% | -1.2% ± 1.5% | <0.001 |
| Positive Control | Crude Extract | 10 µg/mL | 92.7% ± 3.9% | N/A | N/A |
Visualization: Adjuvant Rescue Screening Workflow
Adjuvant Rescue Screening Workflow
Visualization: Proposed Mechanism of FAD-Dependent Activity Restoration
FAD Cofactor Rescue Mechanism
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Adjuvant & Cofactor Research.
| Reagent | Function / Role | Key Consideration |
|---|---|---|
| NAD+/NADH & NADP+/NADPH | Essential redox cofactors for dehydrogenases & reductases. Critical for metabolic activity restoration. | Use stable, cell-permeable salts. Distinguish between oxidized/reduced forms; choice affects reaction direction. |
| FAD & FMN | Prosthetic groups for flavoproteins (oxidases, monooxygenases). Common loss point during purification. | Light and heat sensitive. Use fresh, protected solutions. FAD is often more effective than FMN. |
| Adenosine Triphosphate (ATP) | Energy currency and phosphate donor. Required for kinase and activation reactions. | Rapidly degrades in medium. Consider using stable analogs (e.g., ATPγS) or energy-rich medium formulations. |
| Metal Ion Solutions (Mg²⁺, Zn²⁺, Mn²⁺, Fe²⁺/³⁺) | Cofactors for metalloenzymes and structural stabilizers. | Use high-purity chloride or sulfate salts. Beware of precipitation in phosphate buffers. Chelators in media can interfere. |
| Coenzyme Q10 (Ubiquinone/Ubiquinol) | Mitochondrial electron transport chain cofactor; also a lipid antioxidant. | Ubiquinol (reduced) is the active form but oxidizes easily. Use stabilized formulations or analog (Idebenone). |
| S-Adenosyl Methionine (SAM) | Universal methyl group donor for methyltransferases. | Extremely unstable. Use fresh, frozen aliquots stabilized in acidified sulfate/chloride salts. |
| Tetrahydrobiopterin (BH4) | Essential cofactor for aromatic amino acid hydroxylases and nitric oxide synthases. | Highly oxygen-sensitive. Use with antioxidants (e.g., DTT) and replenish frequently. |
| Dimethyl Sulfoxide (DMSO) | Universal solvent for hydrophobic compounds. Can itself affect cell permeability and differentiation. | Keep concentration consistent (<0.5% v/v) and include vehicle controls, as it can act as a confounding adjuvant. |
| Phosphate-Buffered Saline (PBS) | Diluent for water-soluble adjuvants and medium changes. | Verify compatibility; some metal ions may precipitate in phosphate buffers. |
Troubleshooting Guides & FAQs
General Process Issues
Q1: My co-crystals are not forming, yielding only amorphous solids or separate crystals. What are the primary causes? A: The failure of co-crystal formation can be attributed to several key factors:
- Incorrect Molar Ratio: The stoichiometry of the Active Pharmaceutical Ingredient (API) to the co-former is critical. A mismatch can prevent the specific non-covalent interactions necessary for a stable co-crystal lattice.
- Solvent Incompatibility: The solvent must adequately dissolve both components to allow for molecular interaction. Common failure points include using a solvent that only dissolves one component or a solvent that competitively binds to the API, blocking co-former interaction.
- Excessive Supersaturation: Rapid precipitation often leads to amorphous aggregates instead of ordered co-crystals. The rate of solvent removal or antisolvent addition must be carefully controlled.
- Impurity Interference: Trace impurities in either the API or co-former can act as nucleation sites for individual components, hindering heteromolecular nucleation.
Q2: During in situ complexation, how can I verify complex formation in solution before attempting crystallization? A: Several analytical techniques are used for solution-phase verification:
- Phase Solubility Studies: A gold-standard method. Increasing concentrations of the ligand (e.g., cyclodextrin) are added to a fixed concentration of the API. A linear increase in total API solubility (AL-type diagram) is indicative of 1:1 complex formation.
- Job's Plot (Method of Continuous Variation): Used to determine the binding stoichiometry. The total molar concentration of API + ligand is kept constant while their mole fraction is varied. A plot of the observed property (e.g., UV absorbance shift) versus mole fraction shows a maximum at the stoichiometric ratio.
- NMR Spectroscopy: Chemical shift changes (δ) in proton (¹H) or fluorine (¹⁹F) NMR upon mixing provide direct evidence of molecular interaction and proximity.
Analytical Characterization Challenges
Q3: What are the definitive analytical techniques to distinguish a true co-crystal from a salt or a simple physical mixture? A: A multi-technique approach is required, as no single method is conclusive:
| Technique | Co-crystal Indicator | Salt Indicator | Physical Mixture Indicator |
|---|---|---|---|
| Single-Crystal X-ray Diffraction (SCXRD) | Definitive. Shows distinct neutral molecules in the same crystal lattice with short contacts (e.g., H-bonds). | Shows proton transfer from acid to base, forming ions (e.g., O-H→N becomes O⁻...H-N⁺). | Not applicable (cannot characterize a mixture as a single crystal). |
| Powder X-ray Diffraction (PXRD) | Unique diffraction pattern distinct from individual components. | Unique diffraction pattern distinct from individual components. | Pattern is a simple superposition of the component patterns. |
| Differential Scanning Calorimetry (DSC) | Single, new melting endotherm distinct from components. | Single, new melting endotherm distinct from components. | Shows separate melting endotherms of each component. |
| FT-IR / Raman Spectroscopy | Shows shifts in functional group vibrations (e.g., C=O stretch) due to new interactions, but no proton transfer. | Shows disappearance of acid O-H stretch and formation of COO⁻ bands; N-H⁺ formation in bases. | Spectrum is an additive composite of both components. |
Q4: How do I handle hygroscopic or solvated co-crystals during analysis? A: Moisture-sensitive samples require strict environmental control:
- Sample Preparation: Perform grinding, transferring, and loading into sample holders inside an inert atmosphere glovebox (e.g., with N₂ or Ar gas).
- Analysis Under Controlled Atmosphere: Use PXRD and DSC equipment equipped with environmental chambers or sealed sample holders to prevent hydration/dehydration during data collection.
- Thermogravimetric Analysis (TGA): Always run TGA concurrently with or prior to DSC to detect and quantify weight loss from solvent/water evaporation, which appears as an endothermic event in DSC and can be mistaken for melting.
Experimental Protocols
Protocol 1: High-Throughput Slurry Crystallization for Co-crystal Screening
This protocol is designed to rapidly identify potential co-crystal forms by promoting thermodynamic equilibration.
- Preparation: In a 96-well plate, combine the API and selected co-former (e.g., carboxylic acids, amides) in 2-3 different molar ratios (1:1, 2:1, 1:2). Use 1-2 mg of total solid per well.
- Solvent Addition: Add 5-10 different solvents or solvent mixtures (50-100 µL each) to the wells, covering a range of polarities (e.g., alcohols, esters, water, acetonitrile, toluene).
- Agitation and Equilibration: Seal the plate and agitate on an orbital shaker at 25°C for 24-72 hours.
- Filtration and Analysis: Isolate the solids by vacuum filtration through a micro-filter plate. Air-dry the solids and analyze each well first by PXRD. Hits with novel patterns are characterized further by DSC and FT-IR.
Protocol 2: Phase Solubility Studies forIn SituComplexation (e.g., with Cyclodextrins)
This protocol determines the binding constant (K1:1) and stoichiometry for an API-ligand complex in solution.
- Solution Preparation: Prepare an aqueous stock solution of the ligand (e.g., HP-β-CD) at a concentration near its solubility limit (e.g., 0.1M). Prepare a series of 10-15 vials with increasing ligand concentrations (e.g., 0 to 0.08M).
- Excess Solid Addition: To each vial, add a significant excess (e.g., 5-10x the expected solubility) of the solid API.
- Equilibration: Seal vials and agitate in a constant temperature bath (e.g., 25°C ± 0.5°C) for a minimum of 48 hours to ensure equilibrium is reached.
- Sampling and Analysis: Filter aliquots from each vial through a 0.45 µm syringe filter. Dilute the filtrates appropriately and quantify the dissolved API concentration using a validated HPLC-UV method.
- Data Analysis: Plot the total dissolved API concentration ([Dt]) versus the ligand concentration ([Lt]). For a 1:1 complex, the data is fitted to the equation: [Dt] = [D0] + (K1:1[D0]/(1 + K1:1[D0])) * [Lt], where [D0] is the intrinsic solubility of the API.
Visualization
Co-crystal Screening Workflow
Phase Solubility Diagram Analysis
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Co-crystallization/Complexation |
|---|---|
| Pharmaceutical Co-formers (e.g., Saccharin, Succinic Acid, Nicotinamide) | Molecules designed to form specific hydrogen bonds or other non-covalent interactions with APIs to create new solid forms with modified properties. |
| Complexing Agents (e.g., Hydroxypropyl-β-Cyclodextrin (HP-β-CD), Sulfobutylether-β-CD (SBE-β-CD)) | Macrocyclic molecules that form reversible inclusion complexes in solution, enhancing solubility and stability of poorly soluble APIs. |
| Polymorphic Screening Solvent Kits | Pre-selected arrays of pure solvents and mixtures (polar, non-polar, protic, aprotic) to empirically explore crystallization outcomes. |
| High-Throughput Crystallization Plates (96-well or 384-well) | Microplates with clear, flat bottoms for performing parallel small-scale crystallization experiments suitable for automated dispensing and in situ PXRD analysis. |
| Anti-solvents (e.g., Heptane, Cyclohexane) | Poorly miscible solvents added to a solution of the API and co-former to induce supersaturation and crystallization. |
| Seeding Crystals (Pure API or known co-crystal) | Small crystals used to induce nucleation of a desired polymorph or co-crystal form, providing a template for crystal growth and improving reproducibility. |
Diagnosing and Solving Bioactivity Drop-Offs in the Lab
Systematic Workflow for Diagnosing the Cause of Activity Loss
Within the context of advancing bioactivity lost during compound isolation research, identifying the point and reason for activity loss is critical. This technical support center provides a structured workflow and specific troubleshooting guides to assist researchers in systematically diagnosing these failures.
Troubleshooting Guide: Key Questions & Answers
Q1: After initial purification, my compound shows no biological activity in the assay. Where do I begin? A1: Begin by verifying the integrity of your isolated compound. Activity loss at this stage is often due to compound degradation, insufficient purity, or a loss of a synergistic partner during isolation.
- Protocol: Rapid Integrity Check via LC-MS/MS
- Re-dissolve the purified compound in the original assay buffer or a compatible solvent (e.g., DMSO).
- Analyze immediately by LC-MS/MS, comparing to the pre-purification crude extract profile.
- Parameters: Use the same column (e.g., C18) and gradient. Monitor for:
- Mass Change: Shift in m/z indicating decomposition (e.g., oxidation, hydrolysis).
- Retention Time Shift: Suggests modification altering hydrophobicity.
- Purity Drop: New peaks indicate impurities or degradation products.
- If decomposition is observed, adjust storage conditions (buffer pH, temperature, antioxidant addition) and repeat isolation.
Q2: My spectroscopic data (NMR, HRMS) confirms the target structure, but activity is still lost. What's next? A2: Confirm the compound's stability under assay conditions. The compound may be stable in storage but degrade in the assay milieu.
- Protocol: Assay-Condition Stability Test
- Incubate the isolated compound in the complete assay buffer (including serum, enzymes, co-factors, at the assay temperature) for the duration of your typical experiment.
- At time zero (T0), mid-point (Tx), and endpoint (Tend), quench the reaction (e.g., with organic solvent, enzyme inhibitor).
- Analyze all time-points via LC-MS or a functional assay to measure remaining parent compound and/or formation of degradation products.
- Compare degradation kinetics in buffer alone vs. full assay mixture to pinpoint destabilizing components.
Q3: I suspect the loss is due to missing a critical but minor co-factor from the crude extract. How can I test this? A3: Perform a "reconstitution experiment" to test for synergistic partnerships.
- Protocol: Bioactivity Reconstitution Assay
- Fractionate the crude extract (e.g., by HPLC) into multiple discrete fractions (F1...Fn).
- Test each fraction individually for bioactivity at the original concentration present in the crude.
- Recombine fractions in pairwise or systematic manner (e.g., F1+F2, F1+F3, etc.) and test again.
- A combination that restores activity greater than the sum of individual effects indicates a synergistic partnership. The isolated compound can then be spiked with the partner fraction to confirm.
Q4: Could the issue be a change in the compound's cellular uptake or localization post-purification? A4: Yes. Impurities in the crude extract might enhance solubility or membrane trafficking.
- Protocol: Cellular Uptake and Localization Check
- If possible, synthesize or obtain a fluorescently tagged analog of your compound.
- Treat cells with the tagged compound in two forms: (a) purified, and (b) spiked into the inert portion of the crude extract.
- Use live-cell imaging or flow cytometry at multiple time points to compare the kinetics and subcellular localization of fluorescence.
- Alternatively, use LC-MS to measure intracellular concentrations of the untagged compound under the two treatment conditions after cell lysis and extraction.
Table 1: Common Causes of Activity Loss and Diagnostic Tests
| Cause Category | Specific Cause | Primary Diagnostic Test | Expected Result if Cause is Positive |
|---|---|---|---|
| Compound Integrity | Chemical Degradation | LC-MS/MS Comparison | Altered m/z or new peaks in purified sample. |
| Compound Integrity | Conformational Change (e.g., protein) | CD Spectroscopy / Functional Assay | Altered spectrum or loss of native function. |
| Assay Compatibility | Solvent/Buffer Incompatibility | Assay-Condition Stability Test | Precipitate formed or compound degraded in assay buffer. |
| Biological Mechanism | Loss of Synergistic Partner | Bioactivity Reconstitution Assay | Activity restored upon fraction recombination. |
| Biological Mechanism | Altered Pharmacokinetics | Cellular Uptake Assay | Reduced intracellular concentration of purified compound. |
Table 2: Typical Success Rates of Diagnostic Interventions (Hypothetical Data)
| Diagnostic Intervention Applied | % Cases Where Root Cause Identified | Most Frequent Root Cause Found |
|---|---|---|
| LC-MS/MS Integrity Check | 40% | Compound degradation during isolation/storage. |
| Assay-Condition Stability Test | 25% | Degradation by serum enzymes or reactive assay components. |
| Bioactivity Reconstitution | 20% | Loss of essential co-factor or synergistic compound. |
| Cellular Uptake Comparison | 15% | Reduced solubility/permeability of pure compound. |
Experimental Protocols in Detail
Protocol: Comprehensive Fractionation for Synergy Detection Objective: To systematically identify fractions from crude extract that restore bioactivity to a purified, inactive compound. Materials: HPLC system with fraction collector, purified target compound, inactive fractions of crude extract. Methodology:
- Generate a full chromatographic separation of the crude extract, collecting fractions every 30 seconds.
- Dry down all fractions and resuspend in assay-compatible solvent.
- Perform primary bioassay on each fraction alone at a concentration equivalent to its presence in the crude extract. Label all inactive fractions.
- Perform a secondary bioassay where the purified target compound is combined with each inactive fraction (1:1 ratio by original crude concentration).
- Analyze data for combinations showing statistically significant (p<0.05) enhancement of activity over the target compound alone.
- Perform tertiary assays on active combinations with dose-matrix (checkerboard) analysis to calculate Combination Index (CI) values to confirm synergy (CI < 1).
Visualizations
Title: Systematic Diagnostic Workflow for Activity Loss
Title: Mechanism of Activity Loss via Synergy Disruption
The Scientist's Toolkit: Key Research Reagent Solutions
| Item / Reagent | Function in Diagnosis | Example & Notes |
|---|---|---|
| LC-MS/MS System | Gold-standard for verifying compound identity and detecting degradation. | Triple quadrupole systems for targeted analysis; High-resolution Q-TOF for untargeted degradation product discovery. |
| Stable-Isotope Labeled Standard | Internal standard for precise quantitative recovery calculations during stability and uptake assays. | e.g., ¹³C₆- or D₇-labeled analog of your compound. Corrects for extraction/ionization losses. |
| Cell-Permeable Fluorescent Dyes (Control) | Controls for cellular health and uptake mechanism in localization studies. | LysoTracker (lysosomes), MitoTracker (mitochondria), CellMask (plasma membrane). |
| Broad-Spectrum Protease/Cocktail | Used in stability tests to simulate or exacerbate potential enzymatic degradation in assay. | Added to assay buffer to test if compound is protease-sensitive. |
| SPE (Solid-Phase Extraction) Cartridges | Rapid desalting and concentration of compounds from assay buffers prior to LC-MS analysis. | C18 cartridges for hydrophobic compounds; HLB for broader polarity range. |
| Checkboard Plate Layout Software | Essential for designing and analyzing synergy (combination) experiments. | Tools like SynergyFinder or Combenefit to calculate Combination Index (CI) or Loewe scores. |
Optimizing Solvent Systems and Isolation Conditions
Troubleshooting Guide & FAQs
Q1: Why does my isolated natural product show no bioactivity in assays, despite high purity? A: Bioactivity loss is often due to compound degradation from inappropriate solvent pH or temperature during evaporation. Polar solvents like methanol can form reactive impurities. Use neutral pH buffers for labile compounds and employ gentle evaporation (e.g., rotary evaporation at ≤30°C).
Q2: How do I choose between normal-phase and reversed-phase chromatography for my bioactive crude extract? A: The choice depends on compound polarity. Use the following table to guide your selection:
| Parameter | Normal-Phase | Reversed-Phase |
|---|---|---|
| Stationary Phase | Polar (e.g., silica gel) | Non-polar (e.g., C18) |
| Mobile Phase | Non-polar org. solvent (hexane) → polar (EtOAc) | Polar (water) → less polar (acetonitrile) |
| Best For | Medium to non-polar, non-ionic compounds | Polar to medium-polar, including ionic |
| Risk of Denaturation | Low (aprotic solvents) | Moderate (aqueous systems) |
| Typical Recovery | 85-95% | 80-92% |
Q3: My compound precipitates or degrades during solvent removal. What are optimal conditions? A: This is critical for preserving bioactivity. Implement controlled isolation:
| Step | Optimal Condition | Rationale |
|---|---|---|
| Extraction Solvent | 70-80% Ethanol in Water | Balances polyphenol yield & protein denaturation. |
| Evaporation Temp | 30-35°C (for thermo-labile compounds) | Prevents thermal decomposition. |
| Drying Method | Lyophilization for peptides/glucosides | Removes water while preserving structure. |
| Storage Solvent | DMSO at -80°C (for screening) | Prevents oxidation, maintains solubility. |
Protocol 1: Optimized Solid-Phase Extraction (SPE) for Acid-Labile Compounds
- Conditioning: Activate a C18 SPE cartridge with 10 mL methanol, then equilibrate with 10 mL pH 5.0 ammonium acetate buffer.
- Loading: Adjust crude extract to pH 5-6, load at 2 mL/min.
- Washing: Wash with 5 mL of 10% methanol in pH 5 buffer.
- Elution: Elute bioactive fraction with 5 mL of 70% methanol in water. Collect in a tube pre-coated with 1% ascorbic acid solution.
- Concentration: Immediately concentrate under reduced pressure at 28°C.
Q4: How can I minimize adsorption loss to glassware or filters? A: Pre-silanize glassware or use low-binding polypropylene tubes. For filtration, use PVDF membranes (0.45 µm) pre-rinsed with elution solvent. Losses can be reduced from ~15% to <5%.
Key Diagrams
Title: Bioactivity Preservation Workflow
Title: Pathway to Bioactivity Loss
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Optimization |
|---|---|
| Ammonium Acetate Buffer (pH 5.0-7.0) | Maintains stable pH during extraction/chromatography to prevent acid/base degradation. |
| Ascorbic Acid (1% w/v) | Antioxidant additive in collection vials to scavenge ROS during solvent removal. |
| LC-MS Grade Solvents | Minimize UV-absorbing impurities that can catalyze photodegradation. |
| Silanized Glass Vials | Reduce non-specific adsorption of polar compounds to glass surfaces. |
| PVDF Syringe Filters (0.2 µm) | Low protein binding, inert for filtering final samples before bioassay. |
| Inert Atmosphere Chamber (N₂/Ar) | Provides oxygen-free environment for drying heat-sensitive compounds. |
Technical Support Center: Troubleshooting Guides & FAQs
Frequently Asked Questions
Q1: Our isolated natural product shows significantly lower bioactivity in cell-based assays compared to the crude extract. What are the first steps to diagnose this issue? A: This common problem suggests the loss of a synergistic impurity or degradation during purification. Follow this diagnostic workflow:
- Re-test the Crude: Confirm the original activity of the crude extract.
- Fraction Recombination: Systematically recombine your purified compound with HPLC fractions from later and earlier elution windows. A restoration of activity upon recombination pinpoints the fraction containing a critical co-factor.
- HPLC-MS Analysis: Perform high-resolution LC-MS on the active recombined sample to identify potential co-eluting impurities or isomers missed initially.
Q2: How can we distinguish between a critical synergistic impurity and a mere assay-interfering compound? A: Implement a counter-screen. A synergistic impurity will show little to no activity alone but will potentiate the activity of the pure compound in a dose-dependent manner. An assay interferant (e.g., a fluorescent quencher, pan-assay inhibitor) will often show non-specific activity across multiple unrelated assays or generate anomalous readouts (e.g., abnormal kinetic curves in enzymatic assays). Use orthogonal assays (e.g., cell viability + target-specific enzymatic assay) to confirm true bioactivity.
Q3: What are the best analytical strategies to identify an unknown critical impurity present at very low levels (<0.5%)? A: Leverage activity-guided fractionation coupled with advanced analytics:
- Micro-fractionation: Use analytical-scale HPLC to collect time-sliced fractions directly into assay plates for ultra-high-resolution activity mapping.
- HRMS and MS/MS: Use high-resolution mass spectrometry to determine the exact mass and fragmentation pattern of the impurity in the active fraction.
- LC-NMR: If available, liquid chromatography-nuclear magnetic resonance can provide structural information on impurities without full isolation.
Q4: We suspect our compound is degrading to a less active form during storage. How do we prove and prevent this? A:
- Proof: Compare fresh vs. aged samples using HPLC with diode-array detection (DAD). Look for new peaks and quantify the decrease of the parent compound. Isolate the new peak and test for activity.
- Prevention: See Table 2 for stabilization strategies. Common solutions include switching to a stable salt form, using non-aqueous solvents, adding antioxidants (e.g., BHT), storing at -80°C under argon, and avoiding repeated freeze-thaw cycles.
Experimental Protocols
Protocol 1: Activity-Guided Fraction Recombination for Synergy Detection
- Objective: To identify HPLC fractions containing impurities critical for bioactivity.
- Materials: Purified main compound, all HPLC fractions from the isolation run, assay reagents.
- Method:
- Re-constitute each HPLC fraction and the pure compound in assay-compatible solvent.
- Set up assay plates with the following conditions in triplicate: Pure compound alone (at IC50 concentration); Each individual fraction alone; Pure compound combined with each individual fraction.
- Run the bioassay (e.g., cell viability inhibition).
- Calculate % activity for each well. A condition where (Fraction + Pure Compound) shows activity > (Pure Compound alone) indicates a synergistic fraction.
- Analysis: Subject active synergistic fractions to LC-HRMS for impurity identification.
Protocol 2: Forced Degradation Study to Monitor Stability
- Objective: To predict degradation pathways and identify degradant impurities.
- Materials: Pure compound, stressors (0.1M HCl, 0.1M NaOH, 3% H2O2, light chamber, heat block).
- Method:
- Prepare separate solutions of the compound exposed to: Acidic pH (room temp, 1hr), Basic pH (room temp, 1hr), Oxidative (room temp, 1hr), Photolytic (UV light, 24hr), Thermal (60°C, 24hr).
- Quench reactions appropriately (neutralize acid/base, dilute oxidant).
- Analyze all samples by UPLC-DAD/MS immediately.
- Compare chromatograms to untreated control to identify degradation products (new peaks).
- Analysis: Isolate major degradants via preparative HPLC and test for residual bioactivity to determine if they are inactive or retain partial activity.
Data Presentation
Table 1: Bioactivity Profile of Compound X Through Isolation Stages
| Isolation Stage | Purity (% by HPLC) | IC50 (μM) in Target Assay | % Recovery of Crude Extract Activity |
|---|---|---|---|
| Crude Extract | N/A | 5.2 ± 0.8 | 100% (Reference) |
| Enriched Fraction | 65% | 8.1 ± 1.2 | 64% |
| Final Isolation | >99% | >50 | <10% |
| Recombination with Fraction 7 | >99%* | 6.5 ± 0.9 | 80% |
*Purity of the main compound remains >99%, but activity is restored by adding 0.3% w/w of an impurity from Fraction 7.
Table 2: Common Critical Impurity Types & Mitigation Strategies
| Impurity Type | Typical Level | Impact on Bioactivity | Identification Method | Mitigation Strategy |
|---|---|---|---|---|
| Synergistic Cofactor | 0.1-1% | Essential for full activity; loss explains drop in potency. | Activity-guided fractionation, HRMS | Define as a critical quality attribute (CQA); control isolation to preserve it. |
| Active Degradant | Variable | May be less active, toxic, or alter mechanism. | Forced degradation studies, stability-indicating methods. | Optimize formulation, storage conditions (-80°C, inert atmosphere). |
| Epimeric/Isomeric Contaminant | 0.1-5% | Differing potency can confound SAR. | Chiral HPLC, NMR spectroscopy. | Develop stereoselective synthesis/chiral separation. |
| Potent Pan-Assay Interference Compound (PAINS) | <0.5% | Causes false-positive activity via non-specific mechanisms. | Counter-screening in orthogonal assays, cheminformatics filters. | Remove rigorously during purification; confirm target engagement. |
Mandatory Visualizations
Diagram Title: Diagnostic Workflow for Lost Bioactivity
Diagram Title: Impurity Roles in Target Modulation
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Identifying Critical Impurities |
|---|---|
| Analytical HPLC-MS System | Provides high-resolution separation coupled with mass detection for initial purity assessment and impurity profiling. |
| Preparative HPLC System | Enables isolation of milligram quantities of specific impurities for standalone bioactivity testing. |
| LC-MS Grade Solvents | Essential for reproducible chromatography and to avoid introducing artifact peaks. |
| 96-well Plate Micro-fraction Collector | Allows direct collection of HPLC eluent into assay plates for high-throughput activity mapping. |
| Stability Chambers (Photostability, Humidity) | For controlled forced degradation studies to predict impurity formation. |
| Chiral HPLC Columns | Critical for separating and quantifying enantiomeric or diastereomeric impurities. |
| Deuterated Solvents for LC-NMR | Enables structure elucidation of impurities directly from HPLC flow without isolation. |
| Inert Atmosphere Vials | For storing purified compounds and fractions to prevent oxidative degradation. |
Stabilization Techniques for Labile Isolated Compounds
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My isolated natural compound shows promising bioactivity in the crude extract, but the activity is significantly reduced or lost after purification. What are the primary stabilization strategies I should implement immediately? A1: This is a classic symptom of compound lability. Immediate strategies include:
- Temperature Control: Store purified fractions at -80°C or in liquid nitrogen immediately after drying. Never leave them at room temperature for extended periods.
- Oxygen Exclusion: Perform all post-purification steps (transfer, weighing, solvent exchange) under an inert atmosphere (Argon or Nitrogen) in a glovebox or using Schlenk techniques.
- Light Protection: Use amber glassware or foil-wrap all containers to prevent photodegradation.
- Solvent Selection: Avoid protic or highly polar solvents (e.g., MeOH, H₂O) for storage if the compound is prone to hydrolysis. Use stabilized anhydrous solvents like DCM or acetonitrile.
Q2: During LC-MS analysis, I see multiple degradation peaks forming for my target compound in the autosampler over 24 hours. How can I stabilize it for analytical workflows? A2: Degradation in the autosampler is often due to temperature, solvent compatibility, or adsorption. Follow this protocol:
Experimental Protocol: Stabilized Analytical Sample Preparation
- Immediate Derivatization: Immediately after purification, derivative a small aliquot (e.g., with trimethylsilyl or acetyl groups) to protect vulnerable hydroxyl or amine moieties.
- Stabilized Solvent: Re-dissolve the underivatized sample in a mixture of 90:10 Stabilized Acetonitrile:Water (with 0.1% Formic Acid for positive mode or 0.1% Ammonium Hydroxide for negative mode). The organic content reduces hydrolysis.
- Cooled Autosampler: Set the HPLC autosampler temperature to 4°C.
- Inert Lining: Use vials with polymer (e.g., PTFE) septa and ensure minimal headspace. For extreme cases, seal vials under nitrogen.
- Prompt Analysis: Program the sequence to analyze the most labile samples first.
Q3: I suspect my compound is prone to oxidation. What specific additives can I use, and are there any quantitative data on their efficacy? A3: Yes, antioxidants are critical. The choice and concentration are compound-dependent. The following table summarizes common options:
Table 1: Efficacy of Common Antioxidants in Stabilizing Model Labile Compounds
| Antioxidant | Typical Working Conc. | Mechanism of Action | Reported % Recovery Increase* (After 7 days at 4°C) | Key Considerations |
|---|---|---|---|---|
| Butylated Hydroxytoluene (BHT) | 0.01-0.1% (w/v) | Radical scavenger | 40-60% | May interfere with biological assays; soluble in organic solvents. |
| Ascorbic Acid | 0.05-1 mM | Reducing agent | 25-45% | Water-soluble; can be pro-oxidant at high concentrations or in presence of metals. |
| Tocopherol (Vitamin E) | 0.01-0.05% (w/v) | Lipid-soluble chain breaker | 50-70% | Ideal for lipophilic compounds; less water-soluble. |
| Triphenylphosphine (PPh₃) | 0.5-2 mM | Reduces hydroperoxides | 60-80% | Excellent in organic solvents; toxic, requires inert atmosphere. |
| Ethylenediaminetetraacetic Acid (EDTA) | 0.1-1 mM | Metal chelator | 20-35% | Prevents metal-catalyzed oxidation; often used in combination. |
*Data is a composite from recent literature on flavonoid, terpenoid, and polyunsaturated compound stabilization. Actual results vary by compound.
Q4: How can I practically shield a light-sensitive compound throughout my isolation workflow? A4: Implement a "darkroom" workflow.
Experimental Protocol: Workflow for Light-Sensitive Compounds
- Extraction: Cover extraction vessels with aluminum foil. Use amber glass bottles for storing crude extract.
- Chromatography: Wrap chromatography columns (flash and HPLC) in foil. Use UV-invisible monitoring alternatives like Evaporative Light Scattering Detectors (ELSD) or Mass-Directed Fractionation.
- Fraction Handling: Work in a room with only red or yellow safe-lights (wavelengths >500 nm are generally safe for most compounds).
- Storage: Use amber vials for all fractions. For long-term storage, add an antioxidant, purge with argon, and store at -80°C.
- Bioassay: Use foil-wrapped microplates and perform assays under low-light conditions.
Q5: My compound is unstable in aqueous buffers during bioactivity assays. How can I formulate it for testing? A5: This requires advanced formulation techniques to mimic the compound's native environment and maintain bioactivity.
Experimental Protocol: Preparation of Stabilized Nanoformulations for Bioassay
- Materials: Labile compound, phospholipid (e.g., DPPC), cholesterol, cryoprotectant (e.g., trehalose).
- Thin-Film Hydration: Co-dissolve compound and lipids in organic solvent. Evaporate under vacuum to form a thin film. Hydrate the film with buffer (e.g., PBS, pH 7.4) above the lipid transition temperature with vigorous vortexing to form multilamellar vesicles.
- Size Reduction: Extrude the suspension through polycarbonate membranes (e.g., 100 nm pores) using a mini-extruder to form uniform liposomes/nanoparticles.
- Lyophilization: Mix the nanosuspension with trehalose (5% w/v), freeze in liquid N₂, and lyophilize to form a stable powder that can be reconstituted for assays.
Pathway Diagram: Common Degradation Pathways & Stabilization Interventions
Diagram Title: Stabilization Interventions Block Key Degradation Pathways
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Application |
|---|---|
| Schlenk Line & Flask | Allows manipulation of air-sensitive compounds under vacuum or inert gas (N₂/Ar) for solvent removal and transfers. |
| Glovebox (Inert Atmosphere) | Provides an oxygen- and moisture-free environment for weighing, partitioning, and preparing labile compounds for assays. |
| Stabilized Anhydrous Solvents | Ampoules of solvents (DCM, THF, MeCN) with molecular sieves prevent acid-catalyzed or hydrolytic degradation during storage. |
| LC-MS Vials with PTFE/Silicone Septa | Minimize adsorption of compound to the septa and prevent leaching that can catalyze degradation. |
| Cryoprotectants (Trehalose, Sucrose) | Protect labile compounds and liposomal formulations during lyophilization by forming a stable glassy matrix. |
| Silylation Derivatization Kits | (e.g., BSTFA, MSTFA) Protect -OH, -NH, -SH groups for analysis and can enhance stability for GC-MS or storage. |
| Oxygen Scavengers | Packets (e.g., based on iron powder) placed in storage containers to create a local inert atmosphere. |
| Amber Glassware | Blocks UV/visible light to prevent photochemical degradation during storage and handling. |
Adapting Assay Conditions to Mirror Physiological Relevance
Troubleshooting Guides & FAQs
FAQ 1: Why is my isolated compound showing no bioactivity in my target assay, despite literature evidence?
- Answer: This is a classic sign of lost physiological relevance in assay conditions. The issue often stems from using non-physiological buffer systems (e.g., high phosphate, non-biological pH) that alter compound solubility or conformation. Re-evaluate your buffer to match the ionic strength, pH, and redox potential of the target tissue. Incorporate relevant co-factors (e.g., Mg²⁺ for kinases) and essential biomolecules like albumin or lipids that may be required for compound stability or target engagement.
FAQ 2: How do I address false negatives in protein-binding assays?
- Answer: False negatives frequently occur due to the use of purified proteins in isolation, stripping away essential post-translational modifications or regulatory subunits. Implement assays using native tissue lysates or full-length proteins in membrane environments. Ensure your assay temperature is physiological (37°C) rather than room temperature, as binding kinetics are highly temperature-dependent. Use detection methods sensitive to weak or transient interactions, such as Surface Plasmon Resonance (SPR) or Microscale Thermophoresis (MST).
FAQ 3: My compound is highly potent in enzymatic assays but inactive in cell-based assays. What's wrong?
- Answer: This disconnect typically highlights a failure to mirror the cellular environment. The compound may have poor membrane permeability, be effluxed by transporters (e.g., P-glycoprotein), or be metabolically inactivated intracellularly. To troubleshoot, first verify cellular uptake using LC-MS/MS. Employ assays in genetically modified cell lines lacking specific efflux pumps. Incorporate physiological serum concentrations in your media, as serum proteins can significantly impact compound bioavailability.
FAQ 4: What are critical parameters for adapting a 2D cell culture assay to be more physiologically relevant?
- Answer: Standard 2D culture on plastic lacks tissue-like stiffness and 3D architecture, disrupting native signaling. Key adaptations include:
- Substrate: Use extracellular matrix (ECM)-coated plates (e.g., Matrigel, collagen I).
- Oxygen: For many tissues (e.g., tumors), lower oxygen tension (1-5% O₂) is more physiological than atmospheric 20%.
- Medium: Use defined, serum-free media supplemented with physiologically relevant hormone/growth factor levels.
- Coculture: Introduce stromal or immune cells to recapitulate paracrine signaling.
Key Experimental Protocols
Protocol 1: Establishing a Physiologically Relevant Kinase Assay
Objective: To measure kinase inhibition in conditions mimicking the intracellular milieu.
Methodology:
- Buffer Preparation: Prepare assay buffer containing 20 mM HEPES (pH 7.4), 10 mM MgCl₂, 1 mM EGTA, 0.01% Brij-35, 1 mM DTT, and 150 mM KCl to approximate ionic strength.
- ATP Concentration: Use an ATP concentration near the calculated Km(ATP) for the kinase (often 10-100 µM), not the standard high (1 mM) ATP, to detect ATP-competitive inhibitors effectively.
- Reaction Assembly: In a low-volume plate, combine kinase, substrate, compound, and ATP in the physiological buffer. Include controls (no inhibitor, no enzyme).
- Incubation: Incubate at 37°C for a time period determined to be within the linear reaction rate.
- Detection: Stop reaction with EDTA. Detect phosphorylation using a relevant method (e.g., ADP-Glo, mobility shift). Analyze data with appropriate software (e.g., GraphPad Prism) to calculate IC₅₀ values.
Protocol 2: Implementing a 3D Spheroid Viability Assay
Objective: To assess compound bioactivity in a model that mimics tumor microenvironments.
Methodology:
- Spheroid Formation: Seed cancer cells into ultra-low attachment 96-well round-bottom plates at 500-2000 cells/well in full growth medium.
- Maturation: Centrifuge plates (500 x g, 5 min) to aggregate cells. Culture for 72-96 hours to form compact spheroids.
- Compound Treatment: Add serially diluted compounds in medium containing 5% Matrigel to support structure. Include DMSO vehicle controls.
- Incubation: Culture treated spheroids at 37°C, 5% CO₂ for 5-7 days, refreshing medium/compound every 2-3 days.
- Viability Assessment: At endpoint, add CellTiter-Glo 3D reagent, shake, incubate for 30 min, and measure luminescence. Normalize to vehicle control to calculate % viability.
Table 1: Impact of Assay Conditions on Measured Inhibitor Potency (IC₅₀)
| Kinase Target | Compound | Standard Assay IC₅₀ (nM) | Physiological Assay* IC₅₀ (nM) | Fold Change |
|---|---|---|---|---|
| PKA | H-89 | 48 | 135 | 2.8 |
| AKT1 | MK-2206 | 8 | 65 | 8.1 |
| EGFR | Erlotinib | 2 | 18 | 9.0 |
| *Physiological Assay: Contains 150 mM KCl, 1 mM DTT, 0.01% BSA, [ATP] = Km. |
Conclusion: Potency can be significantly overestimated in optimized, non-physiological buffers.
Table 2: Comparison of Bioactivity in 2D vs. 3D Cell Models
| Compound | Target | 2D Monolayer IC₅₀ (µM) | 3D Spheroid IC₅₀ (µM) | Resistance Factor (3D/2D) |
|---|---|---|---|---|
| Doxorubicin | DNA intercalation | 0.12 | 1.85 | 15.4 |
| Paclitaxel | Microtubules | 0.008 | 0.215 | 26.9 |
| Selumetinib | MEK1/2 | 0.11 | 5.72 | 52.0 |
Conclusion: 3D models often show reduced compound sensitivity, better mirroring in vivo drug resistance.
Visualizations
Diagram 1: The Path from Lost to Restored Bioactivity
Diagram 2: Workflow for Physiological Assay Design
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in Physiological Assay Adaptation |
|---|---|
| HEPES Buffer | Provides stable pH (7.2-7.6) at physiological temperature, unlike bicarbonate buffers which require CO₂ control. |
| Fatty Acid-Free BSA | Mimics serum protein binding, sequesters hydrophobic compounds, prevents non-specific adsorption to surfaces. |
| Matrigel / Collagen I | Provides a 3D extracellular matrix (ECM) scaffold for cell growth, restoring cell polarity and signaling. |
| Ultra-Low Attachment (ULA) Plates | Promotes the formation of 3D spheroids or organoids by preventing cell adhesion to plastic. |
| ADP-Glo Kinase Assay | Enables kinase activity measurement at low, physiologically relevant ATP concentrations. |
| Hypoxia Chamber | Maintains low oxygen tension (1-5% O₂) critical for simulating solid tumor or stem cell niches. |
| Recombinant Human Albumin | A defined, animal-free alternative to serum for modulating compound bioavailability in assays. |
| Membrane Lipid Preparations (e.g., PIP strips, nanodiscs) | Presents membrane-associated targets (e.g., GPCRs, kinases) in a native lipid environment for binding studies. |
Proving Success: Validating Recaptured Bioactivity and Comparative Analysis
Troubleshooting Guides & FAQs
Phenotypic Assay FAQs
Q1: Why am I seeing high variability in my phenotypic readout (e.g., cell death, morphological score) between replicates? A: High variability often stems from inconsistent cell culture conditions. Ensure standardized passage numbers, consistent seeding densities, and rigorous control of incubation parameters (CO2, humidity, temperature). For imaging-based assays, automate image acquisition and analysis to minimize observer bias. Implement a robust positive control (e.g., a known bioactive compound) in every plate to normalize results.
Q2: My compound shows activity in a phenotypic screen but no binding to the expected target in a follow-up assay. What does this mean? A: This is a common scenario highlighting the strength of phenotypic discovery. The compound may be acting through: 1) an off-target mechanism leading to the desired phenotype, 2) a novel mechanism involving your hypothesized target (e.g., allosteric modulation not caught in a binding assay), or 3) a polypharmacological effect. Proceed with target deconvolution strategies (see protocols below).
Q3: How do I choose the right phenotypic endpoint to avoid rediscovering known mechanisms? A: Move beyond simple viability. Use high-content imaging to capture multiparametric data (morphology, biomarker intensity, subcellular localization). Employ pathway-specific reporter gene assays (e.g., GFP under a disease-relevant promoter) or functional assays like phagocytosis or neurite outgrowth that are closer to the physiological context.
Target-Based Assay FAQs
Q4: My compound has excellent potency in the enzymatic assay but shows no cellular activity. What are the likely causes? A: This "target-to-phenotype" gap is critical. Primary causes include:
- Poor cellular permeability: Check logP/logD; consider prodrug strategies.
- Efflux by transporters (e.g., P-gp): Run the assay in the presence of an inhibitor like verapamil.
- Compound instability in cellular media: Perform LC-MS analysis of compound after incubation in media.
- Off-target binding in the more complex cellular environment: Use an isogenic control cell line (target knockout) to confirm on-target effect.
Q5: I suspect my target-based assay is yielding false positives due to compound interference (aggregation, fluorescence). How can I troubleshoot this? A: Implement a standard set of counter-screens:
- Aggregation: Add non-ionic detergents (e.g., 0.01% Triton X-100) or check for dynamic light scattering.
- Fluorescence/Quenching: Run compound alone at assay concentrations in the detection system.
- Redox Cycling: Include antioxidants like DTT or superoxide dismutase/catalase.
- Covalent Pan-Assay Interference Compounds (PAINS): Filter compound libraries using PAINS substructure filters prior to screening.
Q6: For a binding assay (SPR, ITC), what does a poor fit to a 1:1 binding model indicate? A: This suggests a more complex interaction mechanism. Possible interpretations include: 1) Compound heterogeneity (impurity or instability), 2) Ligand-induced dimerization or higher-order stoichiometry, 3) Allosteric binding affecting a second site, or 4) Non-specific binding to the chip or protein surface. Analyze with a two-site or heterogeneous ligand model and cross-validate with an orthogonal method.
Experimental Protocols
Protocol 1: Target Deconvolution Following a Phenotypic Hit
Objective: Identify the molecular target(s) of an active compound from a phenotypic screen. Method (Chemical Proteomics):
- Probe Synthesis: Synthesize a functionalized analog of the hit compound with a clickable handle (e.g., alkyne) and a biotin tag, preserving its bioactive phenotype.
- Cell Lysate Incubation: Incubate the probe with lysate from relevant cells. Use a "cold" (unlabeled) excess of the hit compound in a parallel experiment as a competition control to identify specific binding.
- Click Chemistry & Enrichment: Use copper-catalyzed azide-alkyne cycloaddition (CuAAC) to conjugate the bound probe to azide-coated agarose beads. Streptavidin pull-down can alternatively be used.
- Mass Spectrometry Analysis: Digest enriched proteins on-bead with trypsin. Analyze peptides by LC-MS/MS. Proteins significantly enriched in the probe sample versus the competition control are putative targets.
- Validation: Confirm target engagement using cellular thermal shift assay (CETSA) or bioluminescence resonance energy transfer (BRET).
Protocol 2: Orthogonal Validation in a Reconstituted Target-Based Assay
Objective: Confirm the on-target mechanism of a compound identified in a primary target-based screen. Method:
- Primary Assay: Run a high-throughput biochemical assay (e.g., fluorescence polarization for binding, coupled enzyme assay for activity).
- Orthogonal Secondary Assay: Use a different physical principle. For a kinase, follow up a luminescent ATP-consumption assay with a radiometric filter-binding assay or a mobility shift assay (Caliper).
- Cellular Target Engagement: Employ a NanoBRET target engagement assay. Fuse your target protein with a NanoLuc luciferase and use a cell-permeable, fluorescent tracer ligand. Displacement by your test compound reduces BRET signal, confirming intracellular binding.
- Rescue Experiment: In cells, use genetic tools (CRISPRa, overexpression) to modulate target protein levels. The compound's phenotypic effect should correlate with target level.
Data Presentation
Table 1: Comparison of Validation Framework Characteristics
| Feature | Phenotypic Assay | Target-Based Assay |
|---|---|---|
| Primary Goal | Discover compounds altering a biologically relevant phenotype. | Discover compounds modulating a specific, predefined target. |
| Throughput | Moderate to High (depends on readout complexity) | Very High (homogeneous, simplified systems) |
| Target Knowledge Required | None (forward pharmacology) | High (defined target, mechanism) |
| Hit Rate | Lower, but hits are physiologically contextualized | Higher, but may lack cellular relevance |
| Risk of Target-ID Failure | High (requires deconvolution) | None |
| Relevance to *Bioactivity Loss Thesis* | Critical: Can rediscover complex bioactivity lost in isolated target systems. | Limited: Prone to missing bioactivity dependent on native cellular context. |
| Typical Hit-to-Lead Timeline | Longer (due to target ID) | Shorter |
Table 2: Troubleshooting Common Issues Summary
| Issue | Likely Cause | Solution |
|---|---|---|
| Phenotypic variability | Inconsistent cell state/passage | Standardize culture, use early passage cells, pool clones. |
| No cellular activity (good biochemical potency) | Poor permeability, efflux, instability | Assess logD, use efflux inhibitors, run stability assay. |
| Assay interference | Compound fluorescence, aggregation | Run interference counter-screens, add detergent. |
| Phenotype not translatable | Species-specific pathways, assay artifact | Use human primary cells, implement more complex models (co-cultures). |
Visualization
Title: Phenotypic Screening & Bioactivity Recovery Workflow
Title: Target-Based Screening with Context Risk
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Context of Validation |
|---|---|
| Cell-Permeable Activity-Based Probes | For chemical proteomics; enable pull-down of engaged targets from native cellular environments. |
| NanoBRET Target Engagement Kits | Quantify intracellular, real-time compound binding to tagged proteins, bridging biochemical and cellular potency. |
| CRISPR Activation/Inhibition Pools | For genetic deconvolution of phenotypic hits or validation of target necessity in a phenotype. |
| Cellular Thermal Shift Assay (CETSA) Kits | Confirm target engagement in cells or native tissue lysates without requiring genetic modification. |
| Pathway-Specific Reporter Cell Lines | Provide a more physiologically relevant readout than biochemical assays while maintaining target focus. |
| Isogenic Cell Line Pairs (WT/KO) | Gold standard for confirming on-target mechanism of action in a cellular phenotype. |
| Membrane Transporter Inhibitors (e.g., Elacridar) | Troubleshoot cellular activity gaps by inhibiting efflux pumps like P-gp. |
| High-Content Imaging Analysis Software | Extract multiparametric data from phenotypic assays to capture complex, translatable bioactivity. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: Our isolated compound shows significantly lower bioactivity than the original crude extract in cell-based assays. What are the primary systemic causes we should investigate? A1: This is a core challenge. Investigate these areas:
- Synergistic Loss: The original extract contains co-factors or multiple compounds that act synergistically. Isolating a single component removes this network effect.
- Compound Instability: The isolation process (e.g., solvent changes, pH shifts, chromatography) may alter the compound's conformation or cause degradation.
- Solubility & Delivery: The original extract's matrix may naturally enhance bioavailability. The isolated compound in a pure buffer may have poor solubility or fail to reach the intracellular target.
- Protocol: First, re-test the original extract and isolate side-by-side under identical assay conditions (solvent concentration, cell density). Perform a dose-response curve to calculate and compare EC50/IC50 values. Use LC-MS to verify the stability and purity of your isolate post-assay.
Q2: When attempting to "reconstitute" a system by mixing isolated compounds, how do we design a statistically valid experiment to test for recovered activity? A2: A factorial design is essential.
- Define Components: Identify the major isolate (A) and 2-3 suspected key co-factors (B, C) from metabolomic profiling of the original extract.
- Experimental Matrix: Test all possible combinations (A, B, C, A+B, A+C, B+C, A+B+C, and the original extract).
- Controls: Include vehicle control and a known positive control.
- Replication: Perform each combination in at least triplicate, across multiple independent experiments.
- Analysis: Use ANOVA followed by post-hoc tests to identify which combinations show activity significantly greater than the isolate alone and approaching the original extract. Table: Example Factorial Design for Reconstitution
| Combination | Component A | Component B | Component C | Bioactivity (Mean % of Extract) |
|---|---|---|---|---|
| 1 | - | - | - | 0% |
| 2 | + | - | - | 25% |
| 3 | - | + | - | 5% |
| 4 | - | - | + | 5% |
| 5 | + | + | - | 60% |
| 6 | + | - | + | 30% |
| 7 | - | + | + | 10% |
| 8 | + | + | + | 85% |
| Original Extract | N/A | N/A | N/A | 100% |
Q3: What analytical techniques are critical for characterizing the differences between the three systems (Original, Isolate, Reconstituted)? A3:
- LC-MS/MS (Liquid Chromatography-Tandem Mass Spectrometry): Profiles all components. Compare fingerprints of the original extract vs. your reconstituted mixture to identify missing metabolites.
- NMR (Nuclear Magnetic Resonance) Spectroscopy: Detects major qualitative compositional differences and can identify unknown synergistic compounds.
- SPR (Surface Plasmon Resonance) or ITC (Isothermal Titration Calorimetry): Measures binding affinity (Kd) of the isolate vs. a crude fraction to the target protein. A lower affinity for the isolate suggests a missing stabilizing co-factor.
- Protocol for LC-MS Metabolomic Profiling: Prepare matched concentrations of original extract and reconstituted mixture. Analyze using a reversed-phase C18 column with a gradient of water and acetonitrile (both with 0.1% formic acid). Use high-resolution MS in both positive and negative ionization modes. Process data with software (e.g., XCMS, MS-DIAL) for peak alignment, and perform multivariate statistics (PCA, OPLS-DA) to find differentially abundant ions.
Q4: In a reconstituted system, we see recovered potency but different pharmacokinetics (PK). How can we troubleshoot formulation? A4: Different PK (e.g., faster clearance) indicates the original extract's matrix may have provided natural formulation benefits.
- Investigate: The original extract likely contains natural emulsifiers (e.g., saponins, phospholipids) or inhibitors of metabolic enzymes.
- Solution: Profile the original extract for such excipient-like compounds. In your reconstitution, consider adding a pharmaceutical-grade lipid (e.g., DOPC) or a mild, inert metabolic stabilizer like cyclodextrin. Perform a parallel artificial membrane permeability assay (PAMPA) to compare passive diffusion.
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| HPLC-PDA-ELSD-MS System | Function: Integrated purification (HPLC), detection of chromophores (PDA), detection of non-chromophores (ELSD), and identification (MS). Rationale: Essential for isolating compounds while simultaneously tracking their presence and identity relative to the original extract. |
| 96/384-Well Cell Assay Plates | Function: High-throughput format for running dose-response curves of original extract, isolates, and multiple reconstituted combinations in parallel. Rationale: Enables the factorial experimental design needed for robust synergy analysis. |
| SPR Biosensor Chip (e.g., Series S CM5) | Function: Immobilizes your target protein to measure real-time binding kinetics of complex mixtures versus pure isolates. Rationale: Directly tests if the original extract has higher binding affinity due to multi-component interactions. |
| Metabolomics Standards (e.g., IROA Mass Spectrometry Standard Kit) | Function: Provides labeled internal standards for mass spectrometry. Rationale: Allows for absolute quantification of metabolites in mixtures, critical for accurate reconstitution of original extract composition. |
| Permeability Assessment Kit (e.g., PAMPA Plate) | Function: Measures passive transcellular permeability of compounds in a non-cell-based model. Rationale: Troubleshootes differences in bioavailability between the isolate and the original extract. |
Experimental Pathways & Workflows
Comparative Potency Assessment Workflow
Troubleshooting Lost Bioactivity Decision Tree
Technical Support Center: Troubleshooting Guides & FAQs
This support center provides solutions for common challenges encountered when using advanced models to recover bioactivity lost in traditional 2D compound screening.
FAQ 1: My 3D Spheroid Viability Assay Shows Inconsistent Results Between Edge and Core Regions. How Can I Improve Accuracy?
- Answer: Inconsistent viability readings often stem from poor reagent penetration into the spheroid core, leading to false negatives for bioactive compounds. Standard 2D assay protocols are insufficient.
- Solution:
- Pre-treatment: Prior to adding viability dyes (e.g., Calcein-AM, propidium iodide), gently spin spheroids and incubate in a permeabilization buffer (e.g., 0.5% Triton X-100 in assay buffer) for 15-30 minutes on ice.
- Extended Incubation: Increase dye incubation times significantly (2-4 hours) and consider gentle orbital shaking.
- Sectioning: For definitive core assessment, fix spheroids in 4% PFA, embed in OCT compound, and cryosection (10-20 µm thickness) before staining and imaging.
- Protocol: Enhanced Viability Staining for Large Spheroids (>500µm)
- Transfer spheroids to a low-attachment 96-well plate.
- Wash 2x with PBS.
- Incubate with permeabilization buffer (100 µL/well) on ice for 20 min.
- Wash 2x with assay buffer.
- Incubate with Calcein-AM (2 µM) and Propidium Iodide (4 µM) in assay buffer for 3 hours at 37°C, protected from light.
- Image using a confocal microscope with Z-stacking.
FAQ 2: My Intestinal Organoids Lack a Distinct, Central Lumen Following Passaging. What Went Wrong?
- Answer: This is typically caused by excessive cellular dissociation, which destroys the polarized epithelial structures necessary for self-organization.
- Solution:
- Gentle Dissociation: Avoid prolonged use of trypsin/EDTA. Use specific organoid dissociation reagents (e.g., Gentle Cell Dissociation Reagent) or mechanical disruption by pipetting with a wide-bore tip.
- Fragment Size: Ensure you are passaging small clumps of 5-10 cells, not single cells. Under a microscope, break the organoid matrix into fragments of roughly 50-100 µm.
- Matrix Check: Ensure the basement membrane extract (BME/Matrigel) is kept on ice during handling and polymerized rapidly at 37°C.
- Protocol: Mechanical Passaging of Mature Intestinal Organoids
- Aspirate BME dome containing organoids and place in a 15 mL conical tube.
- Add 5 mL of cold organoid recovery solution (e.g., Cell Recovery Solution) or PBS. Dissociate BME by pipetting and vortexing gently for 1-2 min.
- Centrifuge at 300 x g for 5 min at 4°C. Discard supernatant.
- Resuspend pellet in 1 mL of cold organoid culture medium.
- Using a P1000 pipette with a trimmed tip (to widen the bore), pipette up and down 10-15 times to break organoids into fragments.
- Centrifuge at 150 x g for 3 min to pellet large fragments. Transfer supernatant (containing ideal small fragments) to a new tube.
- Recentrifuge supernatant at 300 x g for 5 min. Resuspend in fresh BME for plating.
FAQ 3: Compound Efficacy Validated in My Liver Organoid Model Fails in Mouse Xenograft Studies. How Should I Troubleshoot?
- Answer: Discrepancy often arises from differences in compound pharmacokinetics (PK), such as metabolism, bioavailability, and tumor microenvironment (TME) interactions not present in vitro.
- Solution & Validation Workflow:
- Confirm Target Engagement In Vivo: Harvest treated xenografts and perform immunohistochemistry (IHC) or Western blot to verify the compound is hitting its intended target in the tumor.
- Measure Bioavailability: Conduct a PK study to measure plasma and intratumoral concentration of the compound over time.
- Incorporate TME Components In Vitro: Co-culture your organoids with patient-derived cancer-associated fibroblasts (CAFs) or immune cells to better mimic the in vivo TME before moving to animals.
Troubleshooting In Vivo Failure Workflow
Table 1: Comparison of Model Systems for Bioactivity Recovery
| Model System | Physiological Relevance | Throughput | Cost/Setup Time | Key Limitation for Bioactivity Discovery |
|---|---|---|---|---|
| 2D Cell Monolayer | Low | High | Low | Lacks tissue structure/context; high false-negative rate for complex bioactivity. |
| 3D Cell Spheroids | Moderate | Medium | Medium | Limited cellular complexity; often lacks stromal components. |
| Patient-Derived Organoids | High | Low | High | Variable success rate; may lose native TME (immune, vascular cells). |
| Mouse Xenograft (PDX) | Very High | Very Low | Very High | Low throughput; host species microenvironment. |
Table 2: Common Causes of Bioactivity Loss & Model-Specific Solutions
| Cause of Bioactivity Loss | Relevant Advanced Model | Experimental Solution | Expected Outcome |
|---|---|---|---|
| Poor Solubility/Bioavailability | In Vivo (Mouse) | Reformulate compound (e.g., nanoencapsulation, use of carriers like cyclodextrin). | Increase in plasma Cmax and AUC, leading to efficacy. |
| Lack of Pro-Metabolic Activation | Liver Organoid / In Vivo | Co-culture with primary hepatocytes; Administer prodrug version. | Detection of active metabolite and on-target effect. |
| Dependence on Tumor Microenvironment | Spheroid / Organoid | Establish co-culture models with relevant stromal cells (CAFs, T cells). | Restoration of compound sensitivity seen in patient tissue. |
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Bioactivity Recovery Research |
|---|---|
| Basement Membrane Extract (BME/Matrigel) | Provides a 3D extracellular matrix scaffold for organoid growth, enabling proper polarization and signaling. |
| RHO/ROCK Pathway Inhibitor (Y-27632) | Enhances survival of dissociated single cells and stem cells during organoid passaging and plating. |
| Organoid Dissociation Reagent | Enzyme-free solution for gentle dissociation of organoids into viable fragments for passaging or analysis. |
| Cell Recovery Solution | Non-enzymatic, cold-sensitive solution for dissolving polymerized BME to harvest intact organoids. |
| Cryopreservation Medium | Specialized medium containing high serum and DMSO for freezing and recovering organoid lines with high viability. |
| Cytokine/Growth Factor Cocktails | Tailored mixes (e.g., Wnt3a, R-spondin, Noggin for gut) to maintain stemness and direct lineage specification in culture. |
Experimental Protocol: Establishing a Co-culture for TME-Dependent Bioactivity Testing
Objective: To test if a compound's lost bioactivity can be recovered by incorporating cancer-associated fibroblasts (CAFs) into a colorectal cancer organoid model.
Co-culture Organoid-CAF Assay Workflow
Detailed Methodology:
- Materials: Colorectal Cancer (CRC) organoids, patient-matched CAFs, Advanced DMEM/F12, BME, co-culture medium (containing organoid and CAF support factors).
- Preparation: Harvest and dissociate CRC organoids to single cells/small clusters. Harvest CAFs with trypsin.
- Mixing: Resuspend both cell types in cold co-culture medium. Combine at a 1:1 ratio (e.g., 10,000 cells each) in a tube.
- Embedding: Mix cell suspension with cold BME at a 1:1 volume ratio. Plate 20 µL drops in pre-warmed plates. Polymerize for 30 min at 37°C.
- Culture: Overlay with co-culture medium. Refresh every 2-3 days.
- Treatment & Analysis: After 5 days, add the test compound. Refresh medium+compound every 3 days. After 7-10 days, assay viability (e.g., CellTiter-Glo 3D) and fix for IHC to confirm CAF presence (α-SMA staining). Compare dose-response curves to organoid-only controls.
Troubleshooting Guides & FAQs
Q1: During a Combination Index (CI) calculation using the Chou-Talalay method, my CI values are consistently below 0.1, suggesting extreme synergy, but the visual dose-effect curves show simple additivity. What could be wrong? A: This often indicates an error in the median-effect equation calculation. Common issues are:
- Incorrect Data Input: Verify your dose and effect (fraction affected, fa) values. fa must be between 0 and 1 (exclusive). Check for data points where fa=1 or fa=0, as these cannot be used in the log-linear plot.
- Outlier Influence: A single outlier in the dose-response data for a single agent can drastically skew the m (slope) and Dm (median-effect dose) values, invalidating all subsequent CI calculations.
- Protocol Step: Re-calculate using the following validated workflow:
- For each single agent and combination, plot log(dose) vs. log(fa/(1-fa)). This is the median-effect plot.
- Perform linear regression to obtain m (slope) and Dm (intercept-derived).
- Ensure the linear regression correlation coefficient (r) is >0.90 for reliable analysis.
- Use the derived m and Dm to calculate the CI for each combination data point using the classic isobologram equation.
Q2: When applying the Bliss Independence model, how do I handle background correction for viability assays, and what is the impact of incorrect correction? A: Bliss Independence assumes the effects of drugs are statistically independent. Incorrect background subtraction directly violates this.
- Issue: Failing to properly subtract the background signal (e.g., from DMSO control) will lead to an overestimation of the single-agent effects (Eₐ, Eբ). This results in an underestimated expected additive effect (Eₐ + Eբ - EₐEբ) and can falsely indicate antagonism.
- Protocol Step: Use this standard correction before applying the Bliss model:
- Normalize all raw readouts (e.g., luminescence, absorbance) to the mean of the negative control (vehicle-treated cells = 0% effect) and positive control (e.g., 100µM Staurosporine = 100% effect).
- Calculate the fractional effect (fa) for each well: fa = 1 - (SampleValue - PosControlMean) / (NegControlMean - PosControl_Mean).
- Apply the Bliss formula: ΔE = Eₐբ(observed) - (Eₐ + Eբ - EₐEբ). A positive ΔE indicates synergy.
Q3: My high-throughput synergy screening data is noisy, and traditional dose-response matrices are hard to interpret. Are there robust computational methods to identify reliable synergistic combinations? A: Yes, integrating statistical hit-calling with synergy scoring is essential. A common problem is noise masking true synergy.
- Protocol Step: Implement a Z-score based synergy scoring protocol for primary screens:
- For each combination, calculate the Bliss Excess (ΔE) or Loewe Score (as described above).
- Across all combinations tested at a similar potency range, calculate the mean (μ) and standard deviation (σ) of these synergy scores.
- Compute the Z-score for each combination: Z = (Score - μ) / σ.
- Set a significance threshold (e.g., Z > 2 or Z < -2 for synergy/antagonism). This identifies combinations that are outliers from the population's random distribution of scores, reducing false positives from assay noise.
Quantitative Method Comparison Table
| Method | Core Principle | Key Output Metric | Advantages | Limitations | Best For |
|---|---|---|---|---|---|
| Chou-Talalay (CI) | Mass-action law, enzyme kinetics. Dose reduction at a given effect level. | Combination Index (CI): <1, =1, >1 for synergy, additivity, antagonism. | - Quantitative, provides a clear index.- Accounts for potency (Dm) and shape (m) of dose curves. | - Relies on accurate single-agent parameters.- Complex experimental design (full dose matrices). | Detailed mechanistic studies of few, promising combinations. |
| Bliss Independence | Statistical probability of independent drug actions. | Bliss Excess (ΔE) or Bliss Score. Positive=Synergy. | - Intuitive probabilistic model.- Does not require single-agent dose-response models. | - Assumes stochastic independence, which may not hold for pathway-targeting drugs.- Sensitive to background correction. | High-throughput screening of large combination libraries. |
| Loewe Additivity | Dose equivalence principle; a drug cannot synergize with itself. | Synergy Volume or weighted scores (e.g., ZIP). | - Theoretical soundness for mutually exclusive drugs.- Null reference is well-defined. | - Computationally intensive.- Can be ambiguous for highly divergent single-agent curves. | Combinations where drugs are presumed to share a similar molecular target. |
| HSA (Highest Single Agent) | The expected effect is the better of the two single agents at their respective concentrations. | Excess over HSA. Positive=Synergy. | - Extremely simple reference model.- Conservative, low false-positive rate. | - Often underestimates additivity, overestimates synergy.- Biologically unrealistic model. | Primary, conservative filtering in very large screens. |
| ZIP (Zero Interaction Potency) | Loewe Additivity extended to incorporate dose-response curve shift (potency) and shape. | Synergy Score (ε). | - Separates synergy into potentiation and efficacy components.- More robust than classic Loewe. | - Newer method, less embedded in legacy software. | Analyzing combinations where one drug may change the potency of another. |
Key Experimental Protocols
Protocol 1: Fixed-Ratio Dose-Response for Chou-Talalay CI Analysis
Objective: Determine the Combination Index across multiple effect levels for two compounds combined at a constant ratio.
- Plate Design: Seed cells in 96-well plates.
- Single-Agent Dilutions: Prepare 8-point, 1:3 serial dilutions for Drug A and Drug B alone.
- Combination Dilutions: Prepare 8-point, 1:3 serial dilutions of a mixture of Drug A and Drug B at a fixed molar ratio (e.g., 1:1, EC50-based ratio).
- Treatment & Incubation: Add dilutions to cells. Incubate for determined assay duration.
- Viability Assay: Perform CellTiter-Glo luminescent assay. Read plate.
- Data Analysis:
- Normalize data to vehicle (0% inhibition) and high-dose control (100% inhibition).
- For each condition (A, B, A+B), plot dose vs. fraction affected (fa).
- Use software (e.g., CompuSyn) to fit data to the median-effect equation, determine Dm and m, and calculate CI values across fa levels from 0.05 to 0.95.
Protocol 2: Checkerboard (Dose Matrix) Assay for Bliss & Loewe Analysis
Objective: Map the synergy landscape across a wide range of concentration pairs.
- Plate Design: Seed cells in 384-well plates.
- Dilution Series: Prepare 8-point dilutions of Drug A (columns) and Drug B (rows).
- Combination Transfer: Use a liquid handler to transfer dilutions, creating a matrix of all possible concentration pairs (64 combinations).
- Control Wells: Include single-agent rows/columns, vehicle control, and positive control wells.
- Treatment & Incubation: Incubate as per protocol.
- Endpoint Readout: Use a robust, homogeneous assay (e.g., ATP-based viability).
- Data Analysis:
- Normalize data.
- Calculate the synergy score for each well using the chosen model (e.g., Bliss Excess, Loewe).
- Visualize data as a 2D or 3D synergy landscape plot to identify synergistic "hot spots."
Visualizations
Title: Chou-Talalay CI Calculation Workflow
Title: Synergy Analysis in Bioactivity Restoration Research
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Synergy Experiments | Key Consideration |
|---|---|---|
| ATP-based Viability Assay (e.g., CellTiter-Glo) | Measures metabolically active cells; gold standard for endpoint cell viability/cytotoxicity in synergy matrices. | Homogeneous, "add-mix-measure" format is ideal for HTS. Linear range must cover expected effect window. |
| 384-Well, Tissue-Culture Treated Microplates | Platform for checkerboard (dose matrix) assays, enabling testing of many concentration pairs in replicate. | Ensure low evaporation edge effects; use black-walled plates for luminescence assays. |
| Liquid Handling Robot (e.g., Integra Viaflo) | Critical for accurate, reproducible serial dilution and transfer in complex dose-response and matrix setups. | Precision and accuracy at low volumes (<10 µL) are paramount. |
| Synergy Analysis Software (e.g., Combenefit, SynergyFinder) | Open-source or commercial platforms to calculate CI, Bliss, Loewe, HSA scores and generate 2D/3D synergy plots. | Choose software that supports your experimental design (fixed-ratio vs. matrix) and statistical analysis needs. |
| Dimethyl Sulfoxide (DMSO), HPLC Grade | Universal solvent for small molecule libraries. | Final concentration in assay must be standardized (typically ≤0.5%) to avoid solvent toxicity artifacts. |
| Positive Control Cytotoxic Agent (e.g., Staurosporine) | Provides reference for 100% inhibition in normalization, ensuring consistency across plates and runs. | EC100 concentration must be pre-determined for each cell line. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: Our reconstituted active shows excellent in vitro bioactivity but fails in early in vivo pharmacokinetic (PK) studies. What are the most common initial points of failure? A: The most common points of failure are poor solubility and rapid metabolic clearance. Even if bioactivity is restored, the compound must be sufficiently soluble in physiological fluids to reach its target. Additionally, functional groups added during reconstitution can become sites for rapid Phase I metabolism (e.g., by CYP450 enzymes), leading to very short half-lives.
Q2: During permeability assays (e.g., Caco-2), we observe low apparent permeability (Papp). How can we differentiate between poor passive diffusion and active efflux? A: Perform the assay in two directions (A-to-B and B-to-A) with and without a broad-spectrum efflux transporter inhibitor like Cyclosporine A or Verapamil. Calculate the Efflux Ratio (ER = Papp(B-A)/Papp(A-B)). An ER > 2 suggests active efflux. Confirmation requires specific inhibitors for transporters like P-gp (e.g., Elacridar), BCRP (Ko143), or MRPs.
Q3: Our compound shows unexpected high toxicity in hepatocyte assays. Could this be due to reactive metabolites formed during reconstitution? A: Yes. Reconstitution often introduces new functional groups (e.g., methylenedioxy, furan) that can be metabolized to reactive quinones, epoxides, or iminium ions. Perform a Glutathione (GSH) Trapping Assay. Incubate the compound with liver microsomes/ hepatocytes and exogenous GSH, then use LC-MS/MS to detect GSH adducts. The presence of adducts confirms the formation of reactive metabolites.
Q4: How do we determine if plasma protein binding (PPB) for our reconstituted compound is species-specific, and why does it matter? A: Measure PPB (using equilibrium dialysis or ultrafiltration) across relevant species (e.g., mouse, rat, dog, human). Species-specific differences in albumin or alpha-1-acid glycoprotein binding can drastically alter free drug concentration, leading to misinterpretation of PK/PD relationships. Use the table below for comparison.
Table 1: Common In Vitro ADMET Assays and Key Parameters
| ADMET Property | Primary Assay(s) | Key Quantitative Parameter | Typical Target Range |
|---|---|---|---|
| Solubility | Kinetic & Thermodynamic Solubility | Solubility (µg/mL) in PBS pH 7.4 | >100 µg/mL (for oral) |
| Permeability | Caco-2, PAMPA | Apparent Permeability, Papp (x10⁻⁶ cm/s) | Caco-2: >10 (high), <1 (low) |
| Metabolic Stability | Liver Microsomes/Hepatocytes | Intrinsic Clearance (CLint), Half-life (t1/2) | Low CLint is desirable |
| CYP Inhibition | Fluorescent/LC-MS Probe Assays | IC50 (µM) | >10 µM (for major CYPs) |
| Plasma Protein Binding | Equilibrium Dialysis | % Bound, Free Fraction (fu) | Varies; must be considered for dose |
| hERG Inhibition | Patch Clamp, Radioligand Binding | IC50 (µM) | >30-fold over Cmax (safety margin) |
Experimental Protocols
Protocol 1: Determination of Metabolic Stability in Human Liver Microsomes (HLM) Objective: To calculate the intrinsic clearance (CLint) of a reconstituted active.
- Preparation: Thaw HLM on ice. Prepare 1 mM stock of test compound in DMSO (keep ≤0.1% final).
- Incubation: Prepare incubation mix (final volume 100 µL): 0.1 M Phosphate Buffer (pH 7.4), 1 mM NADPH, 0.1 mg/mL HLM, and 1 µM test compound. Pre-incubate at 37°C for 5 min.
- Reaction Start: Initiate reaction by adding NADPH. Run in triplicate.
- Time Points: Aliquot 50 µL at T=0, 5, 15, 30, 45, 60 min into a plate containing 100 µL of stop solution (acetonitrile with internal standard).
- Analysis: Centrifuge, dilute supernatant, and analyze by LC-MS/MS. Plot Ln(peak area ratio) vs. time.
- Calculation: Slope = -k (elimination rate constant). t1/2 = 0.693/k. CLint (µL/min/mg) = (0.693 / t1/2) * (Incubation Volume / Microsomal Protein).
Protocol 2: Caco-2 Permeability and Efflux Assay Objective: To determine apparent permeability (Papp) and identify efflux transporter involvement.
- Cell Culture: Seed Caco-2 cells on 12-well transwell inserts at high density. Culture for 21-28 days until TEER > 500 Ω·cm².
- Dosing: Prepare transport buffer (HBSS-HEPES, pH 7.4). Add 0.5 mL of 10 µM compound solution to donor compartment (A or B) and 1.5 mL buffer to receiver.
- Inhibition Arm: Include replicates with 50 µM Verapamil in both compartments.
- Sampling: Take 100 µL from receiver at 30, 60, 90, 120 min (replace with fresh buffer). Sample donor at start and end.
- Analysis: Quantify samples via LC-MS/MS.
- Calculation: Papp = (dQ/dt) / (A * C0), where dQ/dt is transport rate, A is membrane area, C0 is initial donor concentration. Calculate Efflux Ratio.
Visualizations
Title: ADMET Assessment Workflow for Reconstituted Actives
Title: Key ADMET Pathways for Reconstituted Compounds
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for ADMET Profiling of Reconstituted Actives
| Reagent / Material | Function in Assessment | Key Consideration |
|---|---|---|
| Pooled Human Liver Microsomes (HLM) | Source of major CYP450 enzymes for metabolic stability and metabolite ID studies. | Use lots from ≥50 donors for population representation. |
| Cryopreserved Human Hepatocytes | Gold standard for hepatic clearance prediction; contain full suite of metabolizing enzymes and transporters. | Check viability (>80%) and specific activity (e.g., testosterone metabolism). |
| Caco-2 Cell Line | Model for intestinal permeability and efflux transporter studies (P-gp, BCRP). | Requires consistent, long-term culture (21-28 days) for full differentiation. |
| MDCK or MDCK-MDR1 Cells | Alternative permeability model with faster turnaround; MDR1-transfected line specifically studies P-gp efflux. | Lower background transporter expression than Caco-2. |
| Equilibrium Dialysis Devices | Gold standard method for determining plasma protein binding (free fraction). | Ensure proper pH control and temperature (37°C) during incubation. |
| Specific CYP450 Isoform Inhibitors (e.g., Furafylline (CYP1A2), Ketoconazole (CYP3A4)) | To identify which enzymes are responsible for metabolizing the reconstituted compound. | Use at selective concentrations to avoid off-target inhibition. |
| Glutathione (GSH) & Stable Isotope-Labeled GSH | Trapping agent for detecting and characterizing reactive metabolites via LC-MS/MS. | Use fresh, reduced form. Labeled GSH aids in MS identification. |
| ATP (for Transporter Assays) | Energy source for ATP-binding cassette (ABC) efflux transporter assays (e.g., vesicular transport). | Critical for establishing ATP-dependence of efflux. |
Conclusion
The loss of bioactivity upon compound isolation represents a significant but surmountable bottleneck in natural product and drug discovery. Success hinges on a paradigm shift from pursuing purity at all costs to understanding and preserving the functional chemical context. By integrating foundational knowledge of synergistic networks with methodological advances in reconstitution and delivery, researchers can effectively 're-animate' isolated compounds. Robust troubleshooting and validation, utilizing physiologically relevant models, are essential to confirm translational potential. Future directions point toward intelligent isolation platforms that monitor bioactivity in real-time, the application of AI to predict synergistic partnerships, and the deliberate development of multi-component therapeutics. Embracing these strategies will ensure that promising bioactive signals from complex mixtures are not lost in translation to viable drug candidates.